[English] 日本語
 Yorodumi
Yorodumi- PDB-1ju7: NMR Solution Structure of the RNA Hairpin Binding Site for the Hi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ju7 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | NMR Solution Structure of the RNA Hairpin Binding Site for the Histone Stem-loop Binding Protein | ||||||||||||||||||
 Components Components | 5'-R(* Keywords KeywordsRNA / hairpin / tetraloop / 3' stack | Function / homology | : / RNA / RNA (> 10) |  Function and homology information Function and homology informationMethod | SOLUTION NMR / simulated annealing, molecular dynamics |  Authors AuthorsDeJong, E.S. / Marzluff, W.F. / Nikonowicz, E.P. |  Citation Citation Journal: RNA / Year: 2002 Journal: RNA / Year: 2002Title: NMR structure and dynamics of the RNA-binding site for the histone mRNA stem-loop binding protein. Authors: DeJong, E.S. / Marzluff, W.F. / Nikonowicz, E.P. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ju7.cif.gz 1ju7.cif.gz | 104.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ju7.ent.gz pdb1ju7.ent.gz | 84.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ju7.json.gz 1ju7.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1ju7_validation.pdf.gz 1ju7_validation.pdf.gz | 326 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1ju7_full_validation.pdf.gz 1ju7_full_validation.pdf.gz | 386.4 KB | Display | |
| Data in XML |  1ju7_validation.xml.gz 1ju7_validation.xml.gz | 4.7 KB | Display | |
| Data in CIF |  1ju7_validation.cif.gz 1ju7_validation.cif.gz | 6.9 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ju/1ju7 https://data.pdbj.org/pub/pdb/validation_reports/ju/1ju7 ftp://data.pdbj.org/pub/pdb/validation_reports/ju/1ju7 ftp://data.pdbj.org/pub/pdb/validation_reports/ju/1ju7 | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 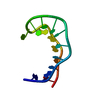
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: RNA chain | Mass: 8928.382 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: This sequence occurs naturally at the 3' end of the replication-dependent histone mRNAs of vertebrates. This sequence corresponds to the mouse H4-12 gene. References: GenBank: 51308 |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||
| NMR details | Text: This structure was determined using a variety of 2D and 3D homo- and hetero-nuclear NMR methods. |
- Sample preparation
Sample preparation
| Details |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions |
| ||||||||||||||||||||
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Bruker AMX / Manufacturer: Bruker / Model: AMX / Field strength: 500 MHz |
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing, molecular dynamics / Software ordinal: 1 Details: Calculations were performed using 232 conformationally restrictive NOE derived distance constraints and 55 backbone and 15 ribose torsion angle constraints. Base pair constraints were ...Details: Calculations were performed using 232 conformationally restrictive NOE derived distance constraints and 55 backbone and 15 ribose torsion angle constraints. Base pair constraints were introduced for six base pairs using heavy atom-heavy atom constraints. Coordinates are for the hairpin domain only. 5' flanking (ggccaaa) and 3' flanking (accca) coordinates are not included with this deposition. Few constraints were obtained for these very dynamic regions and although they were included as part of the structure calculation, their conformations appear to be randomly distributed. | ||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||
| NMR ensemble | Conformer selection criteria: structures with acceptable covalent geometry, structures with the least restraint violations, structures with the lowest energy Conformers calculated total number: 60 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller








 PDBj
PDBj





























 X-PLOR
X-PLOR