+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ih6 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
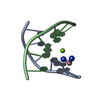
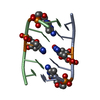
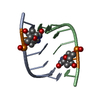
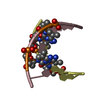
| Title | Multi-Conformation Crystal Structure of GGBr5CGCC | ||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / B to A DNA transition / DNA transition / structural transition | Function / homology | DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 1.45 Å MAD / Resolution: 1.45 Å  Authors AuthorsVargason, J.M. / Henderson, K. / Ho, P.S. |  Citation Citation Journal: Proc.Natl.Acad.Sci.USA / Year: 2001 Journal: Proc.Natl.Acad.Sci.USA / Year: 2001Title: A crystallographic map of the transition from B-DNA to A-DNA. Authors: Vargason, J.M. / Henderson, K. / Ho, P.S. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ih6.cif.gz 1ih6.cif.gz | 69.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ih6.ent.gz pdb1ih6.ent.gz | 53.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ih6.json.gz 1ih6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1ih6_validation.pdf.gz 1ih6_validation.pdf.gz | 391.8 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1ih6_full_validation.pdf.gz 1ih6_full_validation.pdf.gz | 401.5 KB | Display | |
| Data in XML |  1ih6_validation.xml.gz 1ih6_validation.xml.gz | 8.5 KB | Display | |
| Data in CIF |  1ih6_validation.cif.gz 1ih6_validation.cif.gz | 11.6 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ih/1ih6 https://data.pdbj.org/pub/pdb/validation_reports/ih/1ih6 ftp://data.pdbj.org/pub/pdb/validation_reports/ih/1ih6 ftp://data.pdbj.org/pub/pdb/validation_reports/ih/1ih6 | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 1889.101 Da / Num. of mol.: 8 / Source method: obtained synthetically #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.81 Å3/Da / Density % sol: 56.22 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 6 Details: magnesium chloride, sodium cacodylate, spermine tetrahydrochloride, pH 6.0, VAPOR DIFFUSION, SITTING DROP, temperature 298K | ||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 6 | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 105 K | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALS ALS  / Beamline: 5.0.2 / Wavelength: 0.92042, 0.92065, 0.90836 / Beamline: 5.0.2 / Wavelength: 0.92042, 0.92065, 0.90836 | ||||||||||||
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Apr 18, 2000 | ||||||||||||
| Radiation | Monochromator: Si 111 CHANNEL / Protocol: MAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||
| Radiation wavelength |
| ||||||||||||
| Reflection | Resolution: 1.45→20 Å / Num. all: 58509 / Num. obs: 58509 / % possible obs: 99.9 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 10.7 % / Biso Wilson estimate: 20.2 Å2 / Rmerge(I) obs: 0.104 / Net I/σ(I): 22 | ||||||||||||
| Reflection shell | Resolution: 1.45→1.5 Å / Redundancy: 9 % / Rmerge(I) obs: 0.589 / % possible all: 99.6 | ||||||||||||
| Reflection | *PLUS Num. measured all: 628716 / Rmerge(I) obs: 0.11 | ||||||||||||
| Reflection shell | *PLUS % possible obs: 99.6 % / Rmerge(I) obs: 0.668 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 1.45→20 Å / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Chad C. Sines for SHELXL MAD / Resolution: 1.45→20 Å / σ(F): 0 / σ(I): 0 / Stereochemistry target values: Chad C. Sines for SHELXLDetails: This structure was solved using data collected at the peak of Bromine absorption. As a result, the structure factors of the unique reflections and bijvoets have been kept separate for ...Details: This structure was solved using data collected at the peak of Bromine absorption. As a result, the structure factors of the unique reflections and bijvoets have been kept separate for deposition. In addition, the correct calculation for R-factor should take into the account the appropriate scattering factors (f' & f'') at this wavelength.
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.45→20 Å
| ||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 0.9 / Classification: refinement | ||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 20 Å / σ(F): 0 / % reflection Rfree: 10 % / Rfactor obs: 0.138 | ||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller






 PDBj
PDBj


