[English] 日本語
 Yorodumi
Yorodumi- PDB-1i5h: SOLUTION STRUCTURE OF THE RNEDD4 WWIII DOMAIN-RENAL BP2 PEPTIDE C... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1i5h | ||||||
|---|---|---|---|---|---|---|---|
| Title | SOLUTION STRUCTURE OF THE RNEDD4 WWIII DOMAIN-RENAL BP2 PEPTIDE COMPLEX | ||||||
 Components Components |
| ||||||
 Keywords Keywords | LIGASE / Nedd4 / WW domains / ENaC / PY Motif / Liddle syndrome / proline-rich | ||||||
| Function / homology |  Function and homology information Function and homology informationSensory perception of salty taste / Downregulation of ERBB4 signaling / Regulation of PTEN localization / Stimuli-sensing channels / sensory perception of salty taste / ISG15 antiviral mechanism / positive regulation of nucleocytoplasmic transport / Regulation of PTEN stability and activity / Antigen processing: Ubiquitination & Proteasome degradation / neutrophil-mediated killing of bacterium ...Sensory perception of salty taste / Downregulation of ERBB4 signaling / Regulation of PTEN localization / Stimuli-sensing channels / sensory perception of salty taste / ISG15 antiviral mechanism / positive regulation of nucleocytoplasmic transport / Regulation of PTEN stability and activity / Antigen processing: Ubiquitination & Proteasome degradation / neutrophil-mediated killing of bacterium / aldosterone metabolic process / leukocyte activation involved in inflammatory response / cellular response to vasopressin / sensory perception of sour taste / negative regulation of sodium ion transport / cellular response to aldosterone / channel inhibitor activity / nuclear receptor-mediated glucocorticoid signaling pathway / regulation of body fluid levels / response to denervation involved in regulation of muscle adaptation / endocardial cushion development / sodium channel complex / negative regulation of potassium ion export across plasma membrane / epithelial fluid transport / mucus secretion / sodium ion homeostasis / renal system process / regulation protein catabolic process at postsynapse / neutrophil activation involved in immune response / artery smooth muscle contraction / phosphothreonine residue binding / multicellular organismal-level water homeostasis / receptor catabolic process / regulation of protein catabolic process at postsynapse, modulating synaptic transmission / protein targeting to lysosome / ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway / potassium ion homeostasis / HECT-type E3 ubiquitin transferase / intracellular sodium ion homeostasis / proline-rich region binding / potassium channel inhibitor activity / sodium channel inhibitor activity / blood vessel morphogenesis / RNA polymerase binding / sodium ion import across plasma membrane / wound healing, spreading of epidermal cells / beta-2 adrenergic receptor binding / lysosomal transport / negative regulation of vascular endothelial growth factor receptor signaling pathway / lung alveolus development / blood circulation / ligand-gated sodium channel activity / WW domain binding / regulation of postsynaptic neurotransmitter receptor internalization / erythrocyte homeostasis / regulation of dendrite morphogenesis / response to food / sodium channel activity / neuromuscular junction development / regulation of synapse organization / sodium ion transport / microvillus / progesterone receptor signaling pathway / outflow tract morphogenesis / phosphoserine residue binding / protein monoubiquitination / protein K63-linked ubiquitination / ubiquitin ligase complex / postsynaptic cytosol / regulation of sodium ion transport / cellular response to acidic pH / establishment of localization in cell / ionotropic glutamate receptor binding / cytoplasmic vesicle membrane / T cell activation / sodium ion transmembrane transport / ubiquitin binding / regulation of membrane potential / receptor internalization / regulation of blood pressure / multicellular organism growth / neuron projection development / ubiquitin-protein transferase activity / positive regulation of protein catabolic process / cellular response to UV / ubiquitin protein ligase activity / cell cortex / transcription by RNA polymerase II / gene expression / ubiquitin-dependent protein catabolic process / adaptive immune response / transmembrane transporter binding / response to hypoxia / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / apical plasma membrane / immune response / protein ubiquitination / response to xenobiotic stimulus / inflammatory response / external side of plasma membrane Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / torsion angle dynamics | ||||||
 Authors Authors | Kanelis, V. / Rotin, D. / Forman-Kay, J.D. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2001 Journal: Nat.Struct.Biol. / Year: 2001Title: Solution structure of a Nedd4 WW domain-ENaC peptide complex. Authors: Kanelis, V. / Rotin, D. / Forman-Kay, J.D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1i5h.cif.gz 1i5h.cif.gz | 303.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1i5h.ent.gz pdb1i5h.ent.gz | 251.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1i5h.json.gz 1i5h.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/i5/1i5h https://data.pdbj.org/pub/pdb/validation_reports/i5/1i5h ftp://data.pdbj.org/pub/pdb/validation_reports/i5/1i5h ftp://data.pdbj.org/pub/pdb/validation_reports/i5/1i5h | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 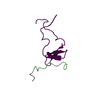
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 5666.237 Da / Num. of mol.: 1 / Fragment: WWIII DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Protein/peptide | Mass: 1725.894 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Description: AN IDENTICAL PEPTIDE WAS MADE SYNTHETICALLY USING STANDARD F-MOC CHEMISTRY FOR NMR SAMPLES REQUIRING UNLABELED BP2 PEPTIDE. Plasmid: PGEX4T2 / Species (production host): Escherichia coli / Production host:  |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 0.01 / pH: 6.5 / Pressure: 1 atm / Temperature: 303 K | |||||||||||||||
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: torsion angle dynamics / Software ordinal: 1 Details: Preliminary structures were calculated using CNS1.0. These structures were then used as input into the program ARIA1.0 for noe assignment and structure refinement. These structures were ...Details: Preliminary structures were calculated using CNS1.0. These structures were then used as input into the program ARIA1.0 for noe assignment and structure refinement. These structures were calculated using 1799 unambiguous and 214 ambiguous NOEs, 14 hydrogen bond restraints, 44 dihedral angle restraints and directly refined against 33 Jhnha coupling constants. | ||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 170 / Conformers submitted total number: 15 |
 Movie
Movie Controller
Controller










 PDBj
PDBj



 NMRPipe
NMRPipe