[English] 日本語
 Yorodumi
Yorodumi- PDB-1epn: A STRUCTURAL COMPARISON OF 21 INHIBITOR COMPLEXES OF THE ASPARTIC... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1epn | ||||||
|---|---|---|---|---|---|---|---|
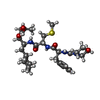
| Title | A STRUCTURAL COMPARISON OF 21 INHIBITOR COMPLEXES OF THE ASPARTIC PROTEINASE FROM ENDOTHIA PARASITICA | ||||||
 Components Components | ENDOTHIAPEPSIN | ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE INHIBITOR / ACID PROTEINASE / HYDROLASE-HYDROLASE INHIBITOR COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Cryphonectria parasitica (chestnut blight fungus) Cryphonectria parasitica (chestnut blight fungus) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.6 Å X-RAY DIFFRACTION / Resolution: 1.6 Å | ||||||
 Authors Authors | Crawford, M. / Cooper, J.B. / Blundell, T.L. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 1994 Journal: Protein Sci. / Year: 1994Title: A structural comparison of 21 inhibitor complexes of the aspartic proteinase from Endothia parasitica. Authors: Bailey, D. / Cooper, J.B. #1:  Journal: J.Mol.Biol. / Year: 1990 Journal: J.Mol.Biol. / Year: 1990Title: The 3D Structure at 2 Angstroms Resolution of Endothiapepsin Authors: Blundell, T.L. / Jenkins, J. / Sewell, B.T. / Pearl, L.H. / Cooper, J.B. / Tickle, I.J. / Veerapandian, B. / Wood, S.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1epn.cif.gz 1epn.cif.gz | 80 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1epn.ent.gz pdb1epn.ent.gz | 56.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1epn.json.gz 1epn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ep/1epn https://data.pdbj.org/pub/pdb/validation_reports/ep/1epn ftp://data.pdbj.org/pub/pdb/validation_reports/ep/1epn ftp://data.pdbj.org/pub/pdb/validation_reports/ep/1epn | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: CIS PROLINE - PRO E 23 2: GLY E 52 - GLN E 53 OMEGA = 211.36 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION 3: CIS PROLINE - PRO E 133 |
- Components
Components
| #1: Protein | Mass: 33813.855 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Cryphonectria parasitica (chestnut blight fungus) Cryphonectria parasitica (chestnut blight fungus)References: UniProt: P11838, endothiapepsin | ||||||
|---|---|---|---|---|---|---|---|
| #2: Chemical | ChemComp-2ZS / | ||||||
| #3: Chemical | | #4: Water | ChemComp-HOH / | Has protein modification | Y | Sequence details | RESIDUES 63A, 80A, 134A, 184A, 203A, 204A, 238A, 282A, 282B, AND 319A ARE INSERTIONS RELATIVE TO ...RESIDUES 63A, 80A, 134A, 184A, 203A, 204A, 238A, 282A, 282B, AND 319A ARE INSERTIONS | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.03 Å3/Da / Density % sol: 39.53 % |
|---|---|
| Crystal grow | *PLUS Method: unknown |
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| Software | Name: RESTRAIN / Classification: refinement | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.6→20 Å / Rfactor obs: 0.17 Details: THE QUANTITY GIVEN IN THE TEMPERATURE FACTOR FIELD OF THE *ATOM* AND *HETATM* RECORDS BELOW IS U**2, WHICH IS THE MEAN-SQUARE AMPLITUDE OF ATOMIC VIBRATION. THE TEMPERATURE FACTOR, B, CAN BE ...Details: THE QUANTITY GIVEN IN THE TEMPERATURE FACTOR FIELD OF THE *ATOM* AND *HETATM* RECORDS BELOW IS U**2, WHICH IS THE MEAN-SQUARE AMPLITUDE OF ATOMIC VIBRATION. THE TEMPERATURE FACTOR, B, CAN BE DERIVED BY THE FOLLOWING RELATION: B = 8 * (PI)**2 * U**2. | ||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→20 Å
| ||||||||||||
| Software | *PLUS Name: RESTRAIN / Classification: refinement | ||||||||||||
| Refinement | *PLUS Rfactor obs: 0.17 | ||||||||||||
| Solvent computation | *PLUS | ||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller

























 PDBj
PDBj