[English] 日本語
 Yorodumi
Yorodumi- PDB-1d6x: THE STRUCTURE OF THE ANTIMICROBIAL PEPTIDE TRITRPTICIN BOUND TO M... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1d6x | ||||||
|---|---|---|---|---|---|---|---|
| Title | THE STRUCTURE OF THE ANTIMICROBIAL PEPTIDE TRITRPTICIN BOUND TO MICELLES-A DISTINCT MEMBRANE-BOUND PEPTIDE FOLD | ||||||
 Components Components | ANTIMICROBIAL PEPTIDE, TRITRPTICIN | ||||||
 Keywords Keywords | IMMUNE SYSTEM / TYPE IV TURN-TYPE III TURN | ||||||
| Method | SOLUTION NMR / RESTRAINED SIMULATED ANNEALING, MOLECULAR DYNAMICS | ||||||
| Model type details | minimized average | ||||||
 Authors Authors | Schibli, D.J. / Hwang, P.M. / Vogel, H.J. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: Structure of the antimicrobial peptide tritrpticin bound to micelles: a distinct membrane-bound peptide fold. Authors: Schibli, D.J. / Hwang, P.M. / Vogel, H.J. #1:  Journal: FEBS Lett. / Year: 1996 Journal: FEBS Lett. / Year: 1996Title: Antimicrobial Activity of a 13 Amino Acid Tryptophan-Rich Peptide Derived From a Putative Porcine Precursor Protein of a Novel Family of Antibacterial Peptides Authors: Lawyer, C. / Pai, S. / Watabe, M. / Borgia, P. / Mashimo, T. / Eagleton, L. / Watabe, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1d6x.cif.gz 1d6x.cif.gz | 96.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1d6x.ent.gz pdb1d6x.ent.gz | 68 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1d6x.json.gz 1d6x.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/d6/1d6x https://data.pdbj.org/pub/pdb/validation_reports/d6/1d6x ftp://data.pdbj.org/pub/pdb/validation_reports/d6/1d6x ftp://data.pdbj.org/pub/pdb/validation_reports/d6/1d6x | HTTPS FTP |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 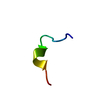
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 1906.286 Da / Num. of mol.: 1 / Fragment: SYNTHETIC CATHELICIDIN FRAGMENT / Source method: obtained synthetically Details: SYNTHETIC PEPTIDE, SEQUENCE FROM A FRAGMENT OF A PORCINE NEUTROPHIL CATHELICIDIN |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||
| NMR details | Text: THIS STRUCTURE WAS DETERMINED USING STANDARD 2D HOMONUCLEAR TECHNIQUES |
- Sample preparation
Sample preparation
| Details |
| ||||||
|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 400 mM SDS / pH: 4.5 / Pressure: AMBIENT / Temperature: 313 K | ||||||
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker AMX / Manufacturer: Bruker / Model: AMX / Field strength: 500 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: RESTRAINED SIMULATED ANNEALING, MOLECULAR DYNAMICS / Software ordinal: 1 Details: THE STRUCTURES ARE BASED ON 163 INTERPROTON RESTRAINTS. OF THESE 47 WERE INTRARESIDUE, 50 SEQUENTIAL, AND 66 SHORT-RANGE | ||||||||||||
| NMR representative | Selection criteria: minimized average structure | ||||||||||||
| NMR ensemble | Conformer selection criteria: structures with favorable non-bond energy Conformers calculated total number: 97 / Conformers submitted total number: 19 |
 Movie
Movie Controller
Controller


 PDBj
PDBj NMRPipe
NMRPipe