[English] 日本語
 Yorodumi
Yorodumi- PDB-1d6b: SOLUTION STRUCTURE OF DEFENSIN-LIKE PEPTIDE-2 (DLP-2) FROM PLATYP... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1d6b | ||||||
|---|---|---|---|---|---|---|---|
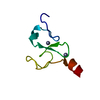
| Title | SOLUTION STRUCTURE OF DEFENSIN-LIKE PEPTIDE-2 (DLP-2) FROM PLATYPUS VENOM | ||||||
 Components Components | DEFENSIN-LIKE PEPTIDE-2 | ||||||
 Keywords Keywords | TOXIN / HELIX / ANTIPARALLEL BETA-SHEET | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / DISTANCE GEOMETRY, SIMULATED ANNEALING, MOLECULAR DYNAMICS, TORSION ANGLE DYNAMICS | ||||||
 Authors Authors | Torres, A.M. / De Plater, G.M. / Doverskog, M. / C Birinyi-Strachan, L. / Nicholson, G.M. / Gallagher, C.H. / Kuchel, P.W. | ||||||
 Citation Citation |  Journal: Biochem.J. / Year: 2000 Journal: Biochem.J. / Year: 2000Title: Defensin-like peptide-2 from platypus venom: member of a class of peptides with a distinct structural fold. Authors: Torres, A.M. / de Plater, G.M. / Doverskog, M. / Birinyi-Strachan, L.C. / Nicholson, G.M. / Gallagher, C.H. / Kuchel, P.W. #1:  Journal: Biochem.J. / Year: 1999 Journal: Biochem.J. / Year: 1999Title: Solution Structure of a Defensin-Like Peptide From Platypus Venom Authors: Torres, A.M. / Wang, X. / Fletcher, J.I. / Alewood, D. / Alewood, P.F. / Smith, R. / Simpson, R.J. / Nicholson, G.M. / Sutherland, S.K. / Gallagher, C.H. / King, G.F. / Kuchel, P.W. #2:  Journal: Biochemistry / Year: 1995 Journal: Biochemistry / Year: 1995Title: Solution Structure of Bovine Neutrophil Beta-Defensin-12: The Peptide Fold of the Beta-Defensins is Identical to That of the Classical Defensins Authors: Zimmermann, G.R. / Legault, P. / Selsted, M.E. / Pardi, A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1d6b.cif.gz 1d6b.cif.gz | 270.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1d6b.ent.gz pdb1d6b.ent.gz | 221.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1d6b.json.gz 1d6b.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/d6/1d6b https://data.pdbj.org/pub/pdb/validation_reports/d6/1d6b ftp://data.pdbj.org/pub/pdb/validation_reports/d6/1d6b ftp://data.pdbj.org/pub/pdb/validation_reports/d6/1d6b | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 5121.899 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|---|
| NMR experiment | Type: 2D NOESY |
| NMR details | Text: THE STRUCTURES WERE DETERMINED USING STANDARD 2D HOMONUCLEAR TECHNIQUES. |
- Sample preparation
Sample preparation
| Details | Contents: 1MM DLP-2 |
|---|---|
| Sample conditions | Ionic strength: NO SALT ADDED / pH: 3.6 / Pressure: AMBIENT / Temperature: 298 K |
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker AVANCE / Manufacturer: Bruker / Model: AVANCE / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DISTANCE GEOMETRY, SIMULATED ANNEALING, MOLECULAR DYNAMICS, TORSION ANGLE DYNAMICS Software ordinal: 1 Details: THE STRUCTURES ARE BASED ON A TOTAL OF 739 RESTRAINTS, 699 ARE NOE-DERIVED DISTANCE RESTRAINTS, 24 DIHEDRAL ANGLE RESTRAINTS, 16 DISTANCE RESTRAINTS FROM HYDROGEN BONDS. | ||||||||||||||||||||
| NMR representative | Selection criteria: lowest energy | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: LOWEST ENERGY / Conformers calculated total number: 4800 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller












 PDBj
PDBj X-PLOR
X-PLOR