+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1by2 | ||||||
|---|---|---|---|---|---|---|---|
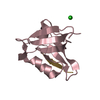
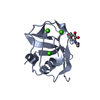
| Title | STRUCTURE OF M2BP SCAVENGER RECEPTOR CYSTEINE-RICH DOMAIN | ||||||
 Components Components | MAC-2 BINDING PROTEIN | ||||||
 Keywords Keywords | EXTRACELLULAR MODULE / SCAVENGER RECEPTOR / TUMOUR-ASSOCIATED ANTIGEN / EXTRACELLULAR MATRIX / GLYCOSYLATED PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationplatelet dense granule lumen / scavenger receptor activity / cellular defense response / Platelet degranulation / extracellular matrix / blood microparticle / cell adhesion / signal transduction / : / extracellular exosome ...platelet dense granule lumen / scavenger receptor activity / cellular defense response / Platelet degranulation / extracellular matrix / blood microparticle / cell adhesion / signal transduction / : / extracellular exosome / extracellular region / membrane Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MIRAS / Resolution: 2 Å MIRAS / Resolution: 2 Å | ||||||
 Authors Authors | Hohenester, E. / Sasaki, T. / Timpl, R. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 1999 Journal: Nat.Struct.Biol. / Year: 1999Title: Crystal structure of a scavenger receptor cysteine-rich domain sheds light on an ancient superfamily. Authors: Hohenester, E. / Sasaki, T. / Timpl, R. #1:  Journal: Embo J. / Year: 1998 Journal: Embo J. / Year: 1998Title: Mac-2 Binding Protein is a Cell-Adhesive Protein of the Extracellular Matrix which Self-Assembles Into Ring-Like Structures and Binds Beta1 Integrins, Collagens and Fibronectin Authors: Sasaki, T. / Brakebusch, C. / Engel, J. / Timpl, R. #2:  Journal: Trends Biochem.Sci. / Year: 1994 Journal: Trends Biochem.Sci. / Year: 1994Title: The Srcr Superfamily: A Family Reminiscent of the Ig Superfamily Authors: Resnick, D. / Pearson, A. / Krieger, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1by2.cif.gz 1by2.cif.gz | 36 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1by2.ent.gz pdb1by2.ent.gz | 22.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1by2.json.gz 1by2.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/by/1by2 https://data.pdbj.org/pub/pdb/validation_reports/by/1by2 ftp://data.pdbj.org/pub/pdb/validation_reports/by/1by2 ftp://data.pdbj.org/pub/pdb/validation_reports/by/1by2 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 12705.093 Da / Num. of mol.: 1 / Fragment: SCAVENGER RECEPTOR CYSTEINE-RICH (SRCR) DOMAIN / Mutation: N-TERMINAL APLA Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Cell line: 293-EBNA / Organ: KIDNEY / Cell line (production host): 293-EBNA / Production host: Homo sapiens (human) / Cell line: 293-EBNA / Organ: KIDNEY / Cell line (production host): 293-EBNA / Production host:  Homo sapiens (human) / References: UniProt: Q08380 Homo sapiens (human) / References: UniProt: Q08380 |
|---|---|
| #2: Sugar | ChemComp-NAG / |
| #3: Water | ChemComp-HOH / |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.6 Å3/Da / Density % sol: 50 % Description: TWO DERIVATIVES (SMCL3 AND UO2SO4) WERE USED AT 3.0 A | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6 / Details: pH 6.0 | |||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7.5 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ELLIOTT GX-21 / Wavelength: 1.5418 ROTATING ANODE / Type: ELLIOTT GX-21 / Wavelength: 1.5418 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Jul 7, 1998 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2→20 Å / Num. obs: 9843 / % possible obs: 99.6 % / Observed criterion σ(I): 0 / Redundancy: 6.2 % / Biso Wilson estimate: 15 Å2 / Rmerge(I) obs: 0.069 / Net I/σ(I): 7.3 |
| Reflection shell | Resolution: 2→2.11 Å / Redundancy: 6 % / Rmerge(I) obs: 0.186 / Mean I/σ(I) obs: 3.7 / % possible all: 100 |
| Reflection | *PLUS Num. measured all: 60557 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIRAS / Resolution: 2→8 Å / Data cutoff high absF: 100000 / Data cutoff low absF: 0 MIRAS / Resolution: 2→8 Å / Data cutoff high absF: 100000 / Data cutoff low absF: 0 Isotropic thermal model: INDIVIDUAL RESTRAINED, RMS BONDS 3.5 A**2 Cross valid method: FREE R-FACTOR / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 20 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati d res low obs: 6 Å / Luzzati sigma a obs: 0.18 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.09 Å / Total num. of bins used: 8
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.84 / Classification: refinement X-PLOR / Version: 3.84 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rwork: 0.27 |
 Movie
Movie Controller
Controller











 PDBj
PDBj






