[English] 日本語
 Yorodumi
Yorodumi- PDB-1bps: MINOR CONFORMER OF A BENZO[A]PYRENE DIOL EPOXIDE ADDUCT OF DA IN ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1bps | ||||||
|---|---|---|---|---|---|---|---|
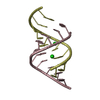
| Title | MINOR CONFORMER OF A BENZO[A]PYRENE DIOL EPOXIDE ADDUCT OF DA IN DUPLEX DNA | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA / DEOXYRIBONUCLEIC ACID / BENZO[A]PYRENE DIOL EPOXIDE ADDUCT / DUPLEX DNA | ||||||
| Function / homology | 1,2,3-TRIHYDROXY-1,2,3,4-TETRAHYDROBENZO[A]PYRENE / DNA Function and homology information Function and homology information | ||||||
| Biological species | synthetic construct (others) | ||||||
| Method | SOLUTION NMR / RELAXATION MATRIX REFINEMENT OF NOE DISTANCE RESTRAINTS, RESTRAINED MOLECULAR DYNAMICS WITH SIMULATED ANNEALING | ||||||
 Authors Authors | Schwartz, J.S. / Rice, J.S. / Luxon, B.A. / Sayer, J.M. / Xie, G. / Yeh, H.J.C. / Liu, X. / Jerina, D.M. / Gorenstein, D.G. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1997 Journal: Biochemistry / Year: 1997Title: Solution structure of the minor conformer of a DNA duplex containing a dG mismatch opposite a benzo[a]pyrene diol epoxide/dA adduct: glycosidic rotation from syn to anti at the modified deoxyadenosine. Authors: Schwartz, J.L. / Rice, J.S. / Luxon, B.A. / Sayer, J.M. / Xie, G. / Yeh, H.J. / Liu, X. / Jerina, D.M. / Gorenstein, D.G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1bps.cif.gz 1bps.cif.gz | 22.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1bps.ent.gz pdb1bps.ent.gz | 14 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1bps.json.gz 1bps.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bp/1bps https://data.pdbj.org/pub/pdb/validation_reports/bp/1bps ftp://data.pdbj.org/pub/pdb/validation_reports/bp/1bps ftp://data.pdbj.org/pub/pdb/validation_reports/bp/1bps | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
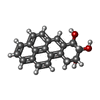
| #1: DNA chain | Mass: 2780.836 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: (+)-(7R,8S,9S,10R)-7,8-DIHYDROXY-9,10-EPOXY-7,8,9,10- TETRAHYDROBENZO[A]PYRENE IS COVALENTLY BONDED TO THE EXOCYCLIC N6 AMINO GROUP OF DEOXYADENOSINE IN THE CENTER OF THE DUPLEX THROUGH ...Details: (+)-(7R,8S,9S,10R)-7,8-DIHYDROXY-9,10-EPOXY-7,8,9,10- TETRAHYDROBENZO[A]PYRENE IS COVALENTLY BONDED TO THE EXOCYCLIC N6 AMINO GROUP OF DEOXYADENOSINE IN THE CENTER OF THE DUPLEX THROUGH TRANS ADDITION AT THE C10 OF THE EPOXIDE. Source: (synth.) synthetic construct (others) |
|---|---|
| #2: DNA chain | Mass: 2716.787 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: BENZO[A]PYRENE DIOL EPOXIDE ADDUCT OF DA / Source: (synth.) synthetic construct (others) |
| #3: Chemical | ChemComp-BAP / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||
| NMR details | Text: MINIMIZED AVERAGE STRUCTURE. THE STRUCTURE WAS DETERMINED USING DISTANCE RESTRAINTS DERIVED FROM 2D NOESY EXPERIMENTS DONE IN 99.999% D2O AND 90% H2O/ 10% D2O. 2D ROESY, TOCSY, AND EXCHANGE- ...Text: MINIMIZED AVERAGE STRUCTURE. THE STRUCTURE WAS DETERMINED USING DISTANCE RESTRAINTS DERIVED FROM 2D NOESY EXPERIMENTS DONE IN 99.999% D2O AND 90% H2O/ 10% D2O. 2D ROESY, TOCSY, AND EXCHANGE-ONLY EXPERIMENTS WERE USED IN THE CHEMICAL SHIFT ASSIGNMENT PROCESS. |
- Sample preparation
Sample preparation
| Sample conditions | Ionic strength: 76 mM / pH: 6.8 / Pressure: 1 atm / Temperature: 288 K |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: RELAXATION MATRIX REFINEMENT OF NOE DISTANCE RESTRAINTS, RESTRAINED MOLECULAR DYNAMICS WITH SIMULATED ANNEALING Software ordinal: 1 Details: THE FOLLOWING RESTRAINTS WERE APPLIED IN THE MOLECULAR DYNAMICS/ENERGY MINIMIZATION CALCULATIONS: (1) INTER-PROTON DISTANCE RESTRAINTS DIRECTLY DERIVED FROM RELAXATION MATRIX ANALYSIS OF NOE ...Details: THE FOLLOWING RESTRAINTS WERE APPLIED IN THE MOLECULAR DYNAMICS/ENERGY MINIMIZATION CALCULATIONS: (1) INTER-PROTON DISTANCE RESTRAINTS DIRECTLY DERIVED FROM RELAXATION MATRIX ANALYSIS OF NOE DATA; (2) IMINO PROTON HYDROGEN BOND RESTRAINTS FOR ALL BASE PAIRS EXCEPT THE CENTRAL DA-DG MISMATCH PAIR. THE MODIFIED DA RESIDUE OF THIS MINOR CONFORMER DISPLAYED A C2'-ENDO SUGAR PUCKER AND AN ANTI GLYCOSIDIC BOND. IT FORMED AN ANTI:ANTI BASE PAIR CONTAINING TWO HYDROGEN BONDS WITH THE OPPOSITE DG. THE MODIFIED DA RESIDUE OF THE MAJOR CONFORMER OF THIS MOLECULE [YEH ET AL. (1995) BIOCHEMISTRY 34, 13570-13581] DISPLAYED C3'-ENDO SUGAR PUCKER AND A SYN GLYCOSIDIC BOND. THE MAJOR CONFORMER MODIFIED DA RESIDUE FORMED A SYN:ANTI BASE PAIR CONTAINING NON-WATSON CRICK HYDROGEN BONDS WITH THE OPPOSITE DG. MORE REFINEMENT DETAILS CAN BE FOUND IN THE PRIMARY JOURNAL CITATION (SCHWARTZ ET AL.,1997). | ||||||||||||||||
| NMR representative | Selection criteria: minimized average structure | ||||||||||||||||
| NMR ensemble | Conformers calculated total number: 1 / Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller





 PDBj
PDBj


 Amber
Amber