[English] 日本語
 Yorodumi
Yorodumi- PDB-1a6b: NMR STRUCTURE OF THE COMPLEX BETWEEN THE ZINC FINGER PROTEIN NCP1... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1a6b | ||||||
|---|---|---|---|---|---|---|---|
| Title | NMR STRUCTURE OF THE COMPLEX BETWEEN THE ZINC FINGER PROTEIN NCP10 OF MOLONEY MURINE LEUKEMIA VIRUS AND A SEQUENCE OF THE PSI-PACKAGING DOMAIN OF HIV-1, 20 STRUCTURES | ||||||
 Components Components |
| ||||||
 Keywords Keywords | Viral protein/DNA / NUCLEOCAPSID PROTEIN / INTERCALATION / NUCLEIC ACID / RETROVIRUS / ZINC FINGER / Viral protein-DNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationhost cell uropod / host cell late endosome membrane / viral budding via host ESCRT complex / host multivesicular body / viral nucleocapsid / structural constituent of virion / host cell plasma membrane / RNA binding / zinc ion binding Similarity search - Function | ||||||
| Method | SOLUTION NMR / DYNAMICAL SIMULATED ANNEALING, ENERGY MINIMIZATION | ||||||
 Authors Authors | Schueler, W. / Dong, C.-Z. / Wecker, K. / Roques, B.P. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: NMR structure of the complex between the zinc finger protein NCp10 of Moloney murine leukemia virus and the single-stranded pentanucleotide d(ACGCC): comparison with HIV-NCp7 complexes. Authors: Schuler, W. / Dong, C. / Wecker, K. / Roques, B.P. #1:  Journal: J.Biomol.NMR / Year: 1994 Journal: J.Biomol.NMR / Year: 1994Title: Three-Dimensional 1H NMR Structure of the Nucleocapsid Protein Ncp10 of Moloney Murine Leukemia Virus Authors: Demene, H. / Jullian, N. / Morellet, N. / De Rocquigny, H. / Cornille, F. / Maigret, B. / Roques, B.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1a6b.cif.gz 1a6b.cif.gz | 320.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1a6b.ent.gz pdb1a6b.ent.gz | 265.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1a6b.json.gz 1a6b.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/a6/1a6b https://data.pdbj.org/pub/pdb/validation_reports/a6/1a6b ftp://data.pdbj.org/pub/pdb/validation_reports/a6/1a6b ftp://data.pdbj.org/pub/pdb/validation_reports/a6/1a6b | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
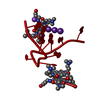
| Deposited unit | 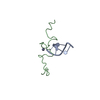
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: DNA chain | Mass: 1465.001 Da / Num. of mol.: 1 / Source method: obtained synthetically |
|---|---|
| #2: Protein/peptide | Mass: 4706.383 Da / Num. of mol.: 1 / Fragment: CENTRAL DOMAIN RESIDUES 14-53 Source method: isolated from a genetically manipulated source Details: ONE ZINC ION BOUND IN CCHC BOX / References: UniProt: P03332 |
| #3: Chemical | ChemComp-ZN / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||
| NMR details | Text: REPRESENTATIVE MODEL NUMBER: ZINC WAS ADDED IN 1.5EQ AS ZNCL2. THE STRUCTURE WAS DETERMINED USING 1H-NMR SPECTROSCOPY ON THE CENTRAL DOMAIN (14-53)NCP10. |
- Sample preparation
Sample preparation
| Sample conditions | pH: 6.5 / Temperature: 293 K |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Bruker AMX600 / Manufacturer: Bruker / Model: AMX600 / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DYNAMICAL SIMULATED ANNEALING, ENERGY MINIMIZATION / Software ordinal: 1 Details: THE PROTEIN PART OF THE STRUCTURE WAS GENERATED IN A SIMULATED ANNEALING CALCULATION AND DOCKED WITH THE NUCLEOTIDE (IDEALIZED CONFORMATION) IN SUCCESSIVE ENERGY MINIMIZATIONS APPLYING FIRST ...Details: THE PROTEIN PART OF THE STRUCTURE WAS GENERATED IN A SIMULATED ANNEALING CALCULATION AND DOCKED WITH THE NUCLEOTIDE (IDEALIZED CONFORMATION) IN SUCCESSIVE ENERGY MINIMIZATIONS APPLYING FIRST INTRA, THEN INTERMOLECULAR NOE RESTRAINTS. THE CONFORMATION OF THE SINGLE STRANDED NUCLEOTIDE GETS DEFINED BY THE CONTACTS TO THE PROTEIN. FIFTY STRUCTURES OF THE COMPLEX WERE GENERATED BY SIMULATED ANNEALING AND ENERGY MINIMIZATION (MAXIMUM GRADIENT O.O2 KCAL/MOL/A2) THE NUCLEOTIDE CONFORMATION WAS KEPT FIXED DURING SIMULATED ANNEALING IN THE CONFORMATION OBTAINED IN THE FIRST STEP. TWENTY STRUCTURES WERE SELECTED WITH RESPECT TO RESTRAINT VIOLATIONS AND TOTAL ENERGY. | ||||||||||||||||
| NMR ensemble | Conformer selection criteria: LEAST RESTRAINT VIOLATIONS, LOWEST TOTAL ENERGY Conformers calculated total number: 50 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller









 PDBj
PDBj


