+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9671 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Adeno-Associated Virus 2 at 2.8 ang | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AAV2 / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont entry into host cell via permeabilization of host membrane / host cell nucleolus / T=1 icosahedral viral capsid / clathrin-dependent endocytosis of virus by host cell / virion attachment to host cell / structural molecule activity Similarity search - Function | |||||||||
| Biological species |  Adeno-associated virus 2 (isolate Srivastava/1982) / Adeno-associated virus 2 (isolate Srivastava/1982) /  Ao-associated virus 2 (isolate Srivastava/1982) Ao-associated virus 2 (isolate Srivastava/1982) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Lou ZY / Zhang R | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2019 Journal: Nat Microbiol / Year: 2019Title: Adeno-associated virus 2 bound to its cellular receptor AAVR. Authors: Ran Zhang / Lin Cao / Mengtian Cui / Zixian Sun / Mingxu Hu / Rouxuan Zhang / William Stuart / Xiaochu Zhao / Zirui Yang / Xueming Li / Yuna Sun / Shentao Li / Wei Ding / Zhiyong Lou / Zihe Rao /   Abstract: Adeno-associated virus (AAV) is a leading vector for virus-based gene therapy. The receptor for AAV (AAVR; also named KIAA0319L) was recently identified, and the precise characterization of AAV-AAVR ...Adeno-associated virus (AAV) is a leading vector for virus-based gene therapy. The receptor for AAV (AAVR; also named KIAA0319L) was recently identified, and the precise characterization of AAV-AAVR recognition is in immediate demand. Taking advantage of a particle-filtering algorithm, we report here the cryo-electron microscopy structure of the AAV2-AAVR complex at 2.8 Å resolution. This structure reveals that of the five Ig-like polycystic kidney disease (PKD) domains in AAVR, PKD2 binds directly to the spike region of the AAV2 capsid adjacent to the icosahedral three-fold axis. Residues in strands B and E, and the BC loop of AAVR PKD2 interact directly with the AAV2 capsid. The interacting residues in the AAV2 capsid are mainly in AAV-featured variable regions. Mutagenesis of the amino acids at the AAV2-AAVR interface reduces binding activity and viral infectivity. Our findings provide insights into the biology of AAV entry with high-resolution details, providing opportunities for the development of new AAV vectors for gene therapy. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9671.map.gz emd_9671.map.gz | 49.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9671-v30.xml emd-9671-v30.xml emd-9671.xml emd-9671.xml | 14.4 KB 14.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9671.png emd_9671.png | 134.8 KB | ||
| Filedesc metadata |  emd-9671.cif.gz emd-9671.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9671 http://ftp.pdbj.org/pub/emdb/structures/EMD-9671 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9671 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9671 | HTTPS FTP |
-Related structure data
| Related structure data |  6ih9MC  9672C  6ihbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9671.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9671.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.932 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
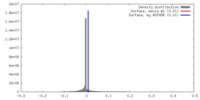
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ao-associated virus 2 (isolate Srivastava/1982)
| Entire | Name:  Ao-associated virus 2 (isolate Srivastava/1982) Ao-associated virus 2 (isolate Srivastava/1982) |
|---|---|
| Components |
|
-Supramolecule #1: Ao-associated virus 2 (isolate Srivastava/1982)
| Supramolecule | Name: Ao-associated virus 2 (isolate Srivastava/1982) / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 648242 Sci species name: Ao-associated virus 2 (isolate Srivastava/1982) Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Virus shell | Shell ID: 1 / Diameter: 280.0 Å |
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Adeno-associated virus 2 (isolate Srivastava/1982) Adeno-associated virus 2 (isolate Srivastava/1982)Strain: isolate Srivastava/1982 |
| Molecular weight | Theoretical: 58.644492 KDa |
| Sequence | String: DGVGNSSGNW HCDSTWMGDR VITTSTRTWA LPTYNNHLYK QISSQSGASN DNHYFGYSTP WGYFDFNRFH CHFSPRDWQR LINNNWGFR PKRLNFKLFN IQVKEVTQND GTTTIANNLT STVQVFTDSE YQLPYVLGSA HQGCLPPFPA DVFMVPQYGY L TLNNGSQA ...String: DGVGNSSGNW HCDSTWMGDR VITTSTRTWA LPTYNNHLYK QISSQSGASN DNHYFGYSTP WGYFDFNRFH CHFSPRDWQR LINNNWGFR PKRLNFKLFN IQVKEVTQND GTTTIANNLT STVQVFTDSE YQLPYVLGSA HQGCLPPFPA DVFMVPQYGY L TLNNGSQA VGRSSFYCLE YFPSQMLRTG NNFTFSYTFE DVPFHSSYAH SQSLDRLMNP LIDQYLYYLS RTNTPSGTTT QS RLQFSQA GASDIRDQSR NWLPGPCYRQ QRVSKTSADN NNSEYSWTGA TKYHLNGRDS LVNPGPAMAS HKDDEEKFFP QSG VLIFGK QGSEKTNVDI EKVMITDEEE IRTTNPVATE QYGSVSTNLQ RGNRQAATAD VNTQGVLPGM VWQDRDVYLQ GPIW AKIPH TDGHFHPSPL MGGFGLKHPP PQILIKNTPV PANPSTTFSA AKFASFITQY STGQVSVEIE WELQKENSKR WNPEI QYTS NYNKSVNVDF TVDTNGVYSE PRPIGTRYLT RNL UniProtKB: Capsid protein VP1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK I |
| Details | The AAV2 particles are purified fron 293T cell, with the yeild of approximately 10^12vg per 10L. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Digitization - Frames/image: 1-19 / Number grids imaged: 1 / Average exposure time: 1.2 sec. / Average electron dose: 1.53 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)