[English] 日本語
 Yorodumi
Yorodumi- EMDB-9260: Electron cryotomogram of Bdellovibrio bacteriovorus acquired by f... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9260 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron cryotomogram of Bdellovibrio bacteriovorus acquired by fast-incremental method | |||||||||
 Map data Map data | Tomographic reconstruction of Bdellovibrio cell. Binned 4 times. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Bdellovibrio bacteriovorus (bacteria) Bdellovibrio bacteriovorus (bacteria) | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Chreifi G / Chen S / Metskas LA / Kaplan M / Jensen GJ | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2019 Journal: J Struct Biol / Year: 2019Title: Rapid tilt-series acquisition for electron cryotomography. Authors: Georges Chreifi / Songye Chen / Lauren Ann Metskas / Mohammed Kaplan / Grant J Jensen /  Abstract: Using a new Titan Krios stage equipped with a single-axis holder, we developed two methods to accelerate the collection of tilt-series. We demonstrate a continuous-tilting method that can record a ...Using a new Titan Krios stage equipped with a single-axis holder, we developed two methods to accelerate the collection of tilt-series. We demonstrate a continuous-tilting method that can record a tilt-series in seconds, but with loss of details finer than ∼4 nm. We also demonstrate a fast-incremental method that can record a tilt-series several-fold faster than current methods and with similar resolution. We characterize the utility of both methods in real biological electron cryotomography workflows. We identify opportunities for further improvements in hardware and software and speculate on the impact such advances could have on structural biology. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9260.map.gz emd_9260.map.gz | 865.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9260-v30.xml emd-9260-v30.xml emd-9260.xml emd-9260.xml | 8.8 KB 8.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9260.png emd_9260.png | 138.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9260 http://ftp.pdbj.org/pub/emdb/structures/EMD-9260 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9260 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9260 | HTTPS FTP |
-Related structure data
| Related structure data |  9261C C: citing same article ( |
|---|---|
| EM raw data |  EMPIAR-10226 (Title: Bdellovibrio bacteriovorus electron cryotomography tilt-series acquired by fast-incremental method EMPIAR-10226 (Title: Bdellovibrio bacteriovorus electron cryotomography tilt-series acquired by fast-incremental methodData size: 5.2 Data #1: Bdellovibrio bacteriovorus electron cryotomography tilt-series acquired by fast-incremental method [tilt series]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9260.map.gz / Format: CCP4 / Size: 1017.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9260.map.gz / Format: CCP4 / Size: 1017.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
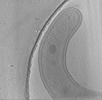
| Annotation | Tomographic reconstruction of Bdellovibrio cell. Binned 4 times. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 10.97 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
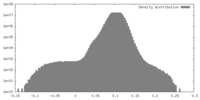
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bdellovibrio bacteriovorus
| Entire | Name:  Bdellovibrio bacteriovorus (bacteria) Bdellovibrio bacteriovorus (bacteria) |
|---|---|
| Components |
|
-Supramolecule #1: Bdellovibrio bacteriovorus
| Supramolecule | Name: Bdellovibrio bacteriovorus / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Bdellovibrio bacteriovorus (bacteria) Bdellovibrio bacteriovorus (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Grid | Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 12.0 nm / Details: unspecified |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 296 K / Instrument: FEI VITROBOT MARK IV |
| Sectioning | Other: NO SECTIONING |
| Fiducial marker | Manufacturer: Ted Pella / Diameter: 10 nm |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: SIMULTANEOUS ITERATIVE (SIRT) / Software - Name: TOMO3D (ver. 2.0) / Number images used: 95 |
|---|---|
| CTF correction | Software - Name:  IMOD (ver. 4.10.18) / Software - details: ctfplotter was used to estimate defocus IMOD (ver. 4.10.18) / Software - details: ctfplotter was used to estimate defocus |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)