[English] 日本語
 Yorodumi
Yorodumi- EMDB-8173: CryoET of MEF Cells and Associated Manual Segmentation of Microtu... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8173 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoET of MEF Cells and Associated Manual Segmentation of Microtubule and Actin | ||||||||||||||||||
 Map data Map data | Mouse Embryonic Fibroblast Cells | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | electron tomography / cryo EM | ||||||||||||||||||
 Authors Authors | Hecksel CW / Darrow MC / Dai W / Galaz-Montoya JG / Chin JA / Mitchell PG / Chen S / Jakana J / Schmid MF / Chiu W | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Microsc Microanal / Year: 2016 Journal: Microsc Microanal / Year: 2016Title: Quantifying Variability of Manual Annotation in Cryo-Electron Tomograms. Authors: Corey W Hecksel / Michele C Darrow / Wei Dai / Jesús G Galaz-Montoya / Jessica A Chin / Patrick G Mitchell / Shurui Chen / Jemba Jakana / Michael F Schmid / Wah Chiu /  Abstract: Although acknowledged to be variable and subjective, manual annotation of cryo-electron tomography data is commonly used to answer structural questions and to create a "ground truth" for evaluation ...Although acknowledged to be variable and subjective, manual annotation of cryo-electron tomography data is commonly used to answer structural questions and to create a "ground truth" for evaluation of automated segmentation algorithms. Validation of such annotation is lacking, but is critical for understanding the reproducibility of manual annotations. Here, we used voxel-based similarity scores for a variety of specimens, ranging in complexity and segmented by several annotators, to quantify the variation among their annotations. In addition, we have identified procedures for merging annotations to reduce variability, thereby increasing the reliability of manual annotation. Based on our analyses, we find that it is necessary to combine multiple manual annotations to increase the confidence level for answering structural questions. We also make recommendations to guide algorithm development for automated annotation of features of interest. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8173.map.gz emd_8173.map.gz | 25.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8173-v30.xml emd-8173-v30.xml emd-8173.xml emd-8173.xml | 14.1 KB 14.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8173.png emd_8173.png | 134.9 KB | ||
| Masks |  emd_8173_msk_1.map emd_8173_msk_1.map emd_8173_msk_10.map emd_8173_msk_10.map emd_8173_msk_11.map emd_8173_msk_11.map emd_8173_msk_12.map emd_8173_msk_12.map emd_8173_msk_13.map emd_8173_msk_13.map emd_8173_msk_14.map emd_8173_msk_14.map emd_8173_msk_2.map emd_8173_msk_2.map emd_8173_msk_3.map emd_8173_msk_3.map emd_8173_msk_4.map emd_8173_msk_4.map emd_8173_msk_5.map emd_8173_msk_5.map emd_8173_msk_6.map emd_8173_msk_6.map emd_8173_msk_7.map emd_8173_msk_7.map emd_8173_msk_8.map emd_8173_msk_8.map emd_8173_msk_9.map emd_8173_msk_9.map | 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB 56.8 MB |  Mask map Mask map | |
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8173 http://ftp.pdbj.org/pub/emdb/structures/EMD-8173 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8173 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8173 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8173.map.gz / Format: CCP4 / Size: 28.4 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_8173.map.gz / Format: CCP4 / Size: 28.4 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
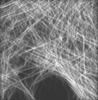
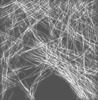
| Annotation | Mouse Embryonic Fibroblast Cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 18.21 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
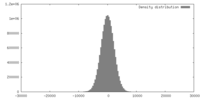
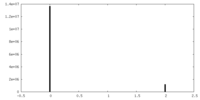
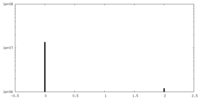
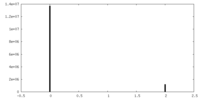
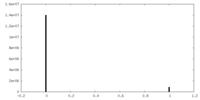
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
+Mask #1
+Mask #10
+Mask #11
+Mask #12
+Mask #13
+Mask #14
+Mask #2
+Mask #3
+Mask #4
+Mask #5
+Mask #6
+Mask #7
+Mask #8
+Mask #9
- Sample components
Sample components
-Entire : Microtubule and Actin from Mouse Embryonic Fibroblast Cells
| Entire | Name: Microtubule and Actin from Mouse Embryonic Fibroblast Cells |
|---|---|
| Components |
|
-Supramolecule #1: Microtubule and Actin from Mouse Embryonic Fibroblast Cells
| Supramolecule | Name: Microtubule and Actin from Mouse Embryonic Fibroblast Cells type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
Details: PBS | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 21 K / Instrument: LEICA EM GP / Details: Plunged into liquid ethane (LEICA EM GP). | |||||||||||||||||||||
| Details | Intact mammalian cells | |||||||||||||||||||||
| Cryo protectant | Ringer's Solution | |||||||||||||||||||||
| Sectioning | Other: NO SECTIONING | |||||||||||||||||||||
| Fiducial marker | Manufacturer: EMS / Diameter: 15 nm |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2100 |
|---|---|
| Temperature | Min: 98.0 K / Max: 98.0 K |
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average exposure time: 1.0 sec. / Average electron dose: 2.6 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 12000 |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Software - Name:  IMOD IMODDetails: Final reconstruction was binned by two before segmentation. Number images used: 27 |
|---|
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)