[English] 日本語
 Yorodumi
Yorodumi- EMDB-75604: The cryo-EM structure of Pakpunavirus P7-1 empty particle neck (p... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The cryo-EM structure of Pakpunavirus P7-1 empty particle neck (portal:H-t-T) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Pakpunavirus P7-1 / Cryo-EM / in situ / structure / neck / portal protein / H-t-T / collar protein / gateway protein / VIRAL PROTEIN / VIRUS | |||||||||
| Biological species |  Pakpunavirus Pakpunavirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Bellis NF / Cingolani G | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: The cryo-EM structure of Pakpunavirus P7-1 neck (portal:H-t-T) Authors: Bellis NF / Cingolani G | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_75604.map.gz emd_75604.map.gz | 62 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-75604-v30.xml emd-75604-v30.xml emd-75604.xml emd-75604.xml | 15.9 KB 15.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_75604_fsc.xml emd_75604_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_75604.png emd_75604.png | 115.1 KB | ||
| Masks |  emd_75604_msk_1.map emd_75604_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-75604.cif.gz emd-75604.cif.gz | 5.4 KB | ||
| Others |  emd_75604_half_map_1.map.gz emd_75604_half_map_1.map.gz emd_75604_half_map_2.map.gz emd_75604_half_map_2.map.gz | 115.8 MB 115.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-75604 http://ftp.pdbj.org/pub/emdb/structures/EMD-75604 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-75604 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-75604 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_75604.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_75604.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.362 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_75604_msk_1.map emd_75604_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
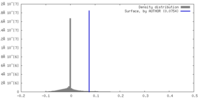
| Density Histograms |
-Half map: #1
| File | emd_75604_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_75604_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Pakpunavirus
| Entire | Name:  Pakpunavirus Pakpunavirus |
|---|---|
| Components |
|
-Supramolecule #1: Pakpunavirus
| Supramolecule | Name: Pakpunavirus / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Pakpunavirus Pakpunavirus |
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pakpunavirus Pakpunavirus |
| Molecular weight | Theoretical: 54.362641 KDa |
| Sequence | String: MAETEKTAPG IPRLRLGEIG STGLKQVNGT ILEERRPELR FPRACRTFQM MAEDPTIKSA LDLFEMMMSR VDWEVDLGVD PDEAMKARG KFLKECMHDM EHSWYSFIKE VTSFYTYGFS VHEMVFKRRE GYPVSKYNDN KFGIKKLPIR SQSTITKWLY S EDGRNFLG ...String: MAETEKTAPG IPRLRLGEIG STGLKQVNGT ILEERRPELR FPRACRTFQM MAEDPTIKSA LDLFEMMMSR VDWEVDLGVD PDEAMKARG KFLKECMHDM EHSWYSFIKE VTSFYTYGFS VHEMVFKRRE GYPVSKYNDN KFGIKKLPIR SQSTITKWLY S EDGRNFLG CEQSLANVVN GDRYVNLGNK DGTVEIPAKK MLLFRVDAKR DSPEGNSPLR ACYNAWRYRV EIEEQESVGV TR DMNGMPT LYLPPRYMSE DATEAERAVY EYYKRVIRNI QMNEQSGLIL PQAFDPESRQ PLFKFELTSS QGSKMYDTDA IIR RWDNKI LQALFADMLK MGQDQVGSYS LAGAKTNIMA MAIESRLREI KDVLDNKLIP TLFALNGDYS PDLPKLQYGE LDEI DLEEF SKGIQRIGSV GGLERDREVY NKIRKALKIK PRPDDEPVDV DNIMGGQSQA GKAGVGNGAS TSAAGRDNAA ANNA |
-Macromolecule #2: Head to Tail protein gp72
| Macromolecule | Name: Head to Tail protein gp72 / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pakpunavirus Pakpunavirus |
| Molecular weight | Theoretical: 18.142607 KDa |
| Sequence | String: MAYTGDPANN PIDRLREIVG DVWEPPMLSD ETYQWVLDKN EGNERRAALE LMRMMLFRLT RGMRERTGDI EVYGAEYFNN YLKALQLIL KDPNIAISLA VPYAGGISKS DMLANDLDPD NVTREFYIGF ARREKLYNQC NPGPQDLSMG CDYLGKLQF |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: OTHER / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)