[English] 日本語
 Yorodumi
Yorodumi- EMDB-7118: Open state structure of the full-length TRPV2 cation channel with... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7118 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Open state structure of the full-length TRPV2 cation channel with a resolved pore turret domain | |||||||||
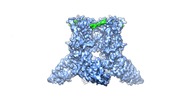
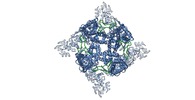
 Map data Map data | WT rat TRPV2 ion channel in an open state and with a resolved pore turret domain. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | channel / TRP / cation / open / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationgrowth cone membrane / TRP channels / response to temperature stimulus / positive regulation of calcium ion import / calcium ion import across plasma membrane / axonal growth cone / positive regulation of axon extension / monoatomic cation channel activity / endomembrane system / calcium channel activity ...growth cone membrane / TRP channels / response to temperature stimulus / positive regulation of calcium ion import / calcium ion import across plasma membrane / axonal growth cone / positive regulation of axon extension / monoatomic cation channel activity / endomembrane system / calcium channel activity / melanosome / lamellipodium / positive regulation of cold-induced thermogenesis / cell body / negative regulation of cell population proliferation / axon / neuronal cell body / cell surface / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Dosey TL / Wang Z | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2019 Journal: Nat Struct Mol Biol / Year: 2019Title: Structures of TRPV2 in distinct conformations provide insight into role of the pore turret. Authors: Timothy L Dosey / Zhao Wang / Guizhen Fan / Zhixian Zhang / Irina I Serysheva / Wah Chiu / Theodore G Wensel /  Abstract: Cation channels of the transient receptor potential (TRP) family serve important physiological roles by opening in response to diverse intra- and extracellular stimuli that regulate their lower or ...Cation channels of the transient receptor potential (TRP) family serve important physiological roles by opening in response to diverse intra- and extracellular stimuli that regulate their lower or upper gates. Despite extensive studies, the mechanism coupling these gates has remained obscure. Previous structures have failed to resolve extracellular loops, known in the TRPV subfamily as 'pore turrets', which are proximal to the upper gates. We established the importance of the pore turret through activity assays and by solving structures of rat TRPV2, both with and without an intact turret at resolutions of 4.0 Å and 3.6 Å, respectively. These structures resolve the full-length pore turret and reveal fully open and partially open states of TRPV2, both with unoccupied vanilloid pockets. Our results suggest a mechanism by which physiological signals, such as lipid binding, can regulate the lower gate and couple to the upper gate through a pore-turret-facilitated mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7118.map.gz emd_7118.map.gz | 14.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7118-v30.xml emd-7118-v30.xml emd-7118.xml emd-7118.xml | 15.8 KB 15.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7118_1.png emd_7118_1.png emd_7118_2.png emd_7118_2.png emd_7118_3.png emd_7118_3.png emd_7118_4.png emd_7118_4.png | 169.2 KB 166.6 KB 171.2 KB 246.9 KB | ||
| Filedesc metadata |  emd-7118.cif.gz emd-7118.cif.gz | 6.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7118 http://ftp.pdbj.org/pub/emdb/structures/EMD-7118 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7118 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7118 | HTTPS FTP |
-Related structure data
| Related structure data |  6bo4MC  7119C  6bo5C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10247 (Title: TRPV2 ion channel gating through allosteric domain coupling revealed by cryo-EM EMPIAR-10247 (Title: TRPV2 ion channel gating through allosteric domain coupling revealed by cryo-EMData size: 3.6 TB Data #1: TRPV2 WT with pore turret domain [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7118.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7118.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
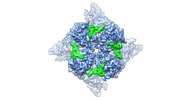
| Annotation | WT rat TRPV2 ion channel in an open state and with a resolved pore turret domain. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.25 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
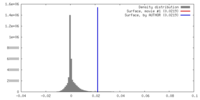
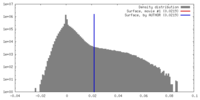
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : TRPV2
| Entire | Name: TRPV2 |
|---|---|
| Components |
|
-Supramolecule #1: TRPV2
| Supramolecule | Name: TRPV2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 320 KDa |
-Macromolecule #1: Transient receptor potential cation channel subfamily V member 2
| Macromolecule | Name: Transient receptor potential cation channel subfamily V member 2 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 79.999422 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVSANGDLHL PISNEQCMPE NNGSLGFEAP TPDRFDRDRL FSVVSRGVPE ELTGLLEYLR WNSKYLTDSA YTEGSTGKTC LMKAVLNLQ DGVNACIMPL LQIDKDSGNP KPLVNAQCTD EFYQGHSALH IAIEKRSLQC VKLLVENGAD VHLRACGRFF Q KHQGTCFY ...String: MVSANGDLHL PISNEQCMPE NNGSLGFEAP TPDRFDRDRL FSVVSRGVPE ELTGLLEYLR WNSKYLTDSA YTEGSTGKTC LMKAVLNLQ DGVNACIMPL LQIDKDSGNP KPLVNAQCTD EFYQGHSALH IAIEKRSLQC VKLLVENGAD VHLRACGRFF Q KHQGTCFY FGELPLSLAA CTKQWDVVTY LLENPHQPAS LEATDSLGNT VLHALVMIAD NSPENSALVI HMYDGLLQMG AR LCPTVQL EEISNHQGLT PLKLAAKEGK IEIFRHILQR EFSGPYQPLS RKFTEWCYGP VRVSLYDLSS VDSWEKNSVL EII AFHCKS PNRHRMVVLE PLNKLLQEKW DRLVSRFFFN FACYLVYMFI FTVVAYHQPS LDQPAIPSSK ATFGESMLLL GHIL ILLGG IYLLLGQLWY FWRRRLFIWI SFMDSYFEIL FLLQALLTVL SQVLRFMETE WYLPLLVLSL VLGWLNLLYY TRGFQ HTGI YSVMIQKVIL RDLLRFLLVY LVFLFGFAVA LVSLSREARS PKAPEDNNST VTEQPTVGQE EEPAPYRSIL DASLEL FKF TIGMGELAFQ EQLRFRGVVL LLLLAYVLLT YVLLLNMLIA LMSETVNHVA DNSWSIWKLQ KAISVLEMEN GYWWCRR KK HREGRLLKVG TRGDGTPDER WCFRVEEVNW AAWEKTLPTL SEDPTETSQV APASTOP UniProtKB: Transient receptor potential cation channel subfamily V member 2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | .5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 98 % / Chamber temperature: 298 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average exposure time: 10.0 sec. / Average electron dose: 63.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)