+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6907 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Atomic structure of the herpes simplex virus type 2 B-capsid | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | herpesvirus / capsid / cryo-em / atomic structure / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationT=16 icosahedral viral capsid / viral capsid assembly / viral process / viral capsid / host cell nucleus / structural molecule activity / DNA binding Similarity search - Function | |||||||||
| Biological species |   Human herpesvirus 2 Human herpesvirus 2 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Yuan S / Wang JL / Zhu DJ / Wang N / Gao Q / Chen WY / Tang H / Wang JZ / Zhang XZ / Liu HR ...Yuan S / Wang JL / Zhu DJ / Wang N / Gao Q / Chen WY / Tang H / Wang JZ / Zhang XZ / Liu HR / Rao ZH / Wang XX | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2018 Journal: Nat Commun / Year: 2018Title: Pushing the resolution limit by correcting the Ewald sphere effect in single-particle Cryo-EM reconstructions. Authors: Dongjie Zhu / Xiangxi Wang / Qianglin Fang / James L Van Etten / Michael G Rossmann / Zihe Rao / Xinzheng Zhang /   Abstract: The Ewald sphere effect is generally neglected when using the Central Projection Theorem for cryo electron microscopy single-particle reconstructions. This can reduce the resolution of a ...The Ewald sphere effect is generally neglected when using the Central Projection Theorem for cryo electron microscopy single-particle reconstructions. This can reduce the resolution of a reconstruction. Here we estimate the attainable resolution and report a "block-based" reconstruction method for extending the resolution limit. We find the Ewald sphere effect limits the resolution of large objects, especially large viruses. After processing two real datasets of large viruses, we show that our procedure can extend the resolution for both datasets and can accommodate the flexibility associated with large protein complexes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6907.map.gz emd_6907.map.gz | 1.2 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6907-v30.xml emd-6907-v30.xml emd-6907.xml emd-6907.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6907.png emd_6907.png | 297.7 KB | ||
| Filedesc metadata |  emd-6907.cif.gz emd-6907.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6907 http://ftp.pdbj.org/pub/emdb/structures/EMD-6907 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6907 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6907 | HTTPS FTP |
-Related structure data
| Related structure data |  5zapMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6907.map.gz / Format: CCP4 / Size: 6.4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6907.map.gz / Format: CCP4 / Size: 6.4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.38 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
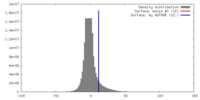
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human herpesvirus 2
| Entire | Name:   Human herpesvirus 2 Human herpesvirus 2 |
|---|---|
| Components |
|
-Supramolecule #1: Human herpesvirus 2
| Supramolecule | Name: Human herpesvirus 2 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1, #4, #2-#3 / NCBI-ID: 10310 / Sci species name: Human herpesvirus 2 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:   Human herpesvirus 2 Human herpesvirus 2 |
-Macromolecule #1: Major capsid protein
| Macromolecule | Name: Major capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human herpesvirus 2 Human herpesvirus 2 |
| Molecular weight | Theoretical: 149.370359 KDa |
| Sequence | String: MAAPARDPPG YRYAAAMVPT GSILSTIEVA SHRRLFDFFA RVRSDENSLY DVEFDALLGS YCNTLSLVRF LELGLSVACV CTKFPELAY MNEGRVQFEV HQPLIARDGP HPVEQPVHNY MTKVIDRRAL NAAFSLATEA IALLTGEALD GTGISLHRQL R AIQQLARN ...String: MAAPARDPPG YRYAAAMVPT GSILSTIEVA SHRRLFDFFA RVRSDENSLY DVEFDALLGS YCNTLSLVRF LELGLSVACV CTKFPELAY MNEGRVQFEV HQPLIARDGP HPVEQPVHNY MTKVIDRRAL NAAFSLATEA IALLTGEALD GTGISLHRQL R AIQQLARN VQAVLGAFER GTADQMLHVL LEKAPPLALL LPMQRYLDNG RLATRVARAT LVAELKRSFC DTSFFLGKAG HR REAIEAW LVDLTTATQP SVAVPRLTHA DTRGRPVDGV LVTTAAIKQR LLQSFLKVED TEADVPVTYG EMVLNGANLV TAL VMGKAV RSLDDVGRHL LDMQEEQLEA NRETLDELES APQTTRVRAD LVAIGDRLVF LEALEKRIYA ATNVPYPLVG AMDL TFVLP LGLFNPAMER FAAHAGDLVP APGHPEPRAF PPRQLFFWGK DHQVLRLSME NAVGTVCHPS LMNIDAAVGG VNHDP VEAA NPYGAYVAAP AGPGADMQQR FLNAWRQRLA HGRVRWVAEC QMTAEQFMQP DNANLALELH PAFDFFAGVA DVELPG GEV PPAGPGAIQA TWRVVNGNLP LALCPVAFRD ARGLELGVGR HAMAPATIAA VRGAFEDRSY PAVFYLLQAA IHGSEHV FC ALARLVTQCI TSYWNNTRCA AFVNDYSLVS YIVTYLGGDL PEECMAVYRD LVAHVEALAQ LVDDFTLPGP ELGGQAQA E LNHLMRDPAL LPPLVWDCDG LMRHAALDRH RDCRIDAGGH EPVYAAACNV ATADFNRNDG RLLHNTQARA ADAADDRPH RPADWTVHHK IYYYVLVPAF SRGRCCTAGV RFDRVYATLQ NMVVPEIAPG EECPSDPVTD PAHPLHPANL VANTVNAMFH NGRVVVDGP AMLTLQVLAH NMAERTTALL CSAAPDAGAN TASTANMRIF DGALHAGVLL MAPQHLDHTI QNGEYFYVLP V HALFAGAD HVANAPNFPP ALRDLARHVP LVPPALGANY FSSIRQPVVQ HARESAAGEN ALTYALMAGY FKMSPVALYH QL KTGLHPG FGFTVVRQDR FVTENVLFSE RASEAYFLGQ LQVARHETGG GVNFTLTQPR GNVDLGVGYT AVAATATVRN PVT DMGNLP QNFYLGRGAP PLLDNAAAVY LRNAVVAGNR LGPAQPLPVF GCAQVPRRAG MDHGQDAVCE FIATPVATDI NYFR RPCNP RGRAAGGVYA GDKEGDVIAL MYDHGQSDPA RPFAATANPW ASQRFSYGDL LYNGAYHLNG ASPVLSPCFK FFTAA DITA KHRCLERLIV ETGSAVSTAT AASDVQFKRP PGCRELVEDP CGLFQEAYPI TCASDPALLR SARDGEAHAR ETHFTQ YLI YDASPLKGLS L UniProtKB: Major capsid protein |
-Macromolecule #2: Triplex capsid protein 2
| Macromolecule | Name: Triplex capsid protein 2 / type: protein_or_peptide / ID: 2 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human herpesvirus 2 Human herpesvirus 2 |
| Molecular weight | Theoretical: 34.373785 KDa |
| Sequence | String: MITDCFEADI AIPSGISRPD AAALQRCEGR VVFLPTIRRQ LALADVAHES FVSGGVSPDT LGLLLAYRRR FPAVITRVLP TRIVACPVD LGLTHAGTVN LRNTSPVDLC NGDPVSLVPP VFEGQATDVR LESLDLTLRF PVPLPTPLAR EIVARLVARG I RDLNPDPR ...String: MITDCFEADI AIPSGISRPD AAALQRCEGR VVFLPTIRRQ LALADVAHES FVSGGVSPDT LGLLLAYRRR FPAVITRVLP TRIVACPVD LGLTHAGTVN LRNTSPVDLC NGDPVSLVPP VFEGQATDVR LESLDLTLRF PVPLPTPLAR EIVARLVARG I RDLNPDPR TPGELPDLNV LYYNGARLSL VADVQQLASV NTELRSLVLN MVYSITEGTT LILTLIPRLL ALSAQDGYVN AL LQMQSVT REAAQLIHPE APMLMQDGER RLPLYEALVA WLAHAGQLGD ILALAPAVRV CTFDGAAVVQ SGDMAPVIRY P UniProtKB: Capsid triplex subunit 2 |
-Macromolecule #3: Triplex capsid protein 1
| Macromolecule | Name: Triplex capsid protein 1 / type: protein_or_peptide / ID: 3 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human herpesvirus 2 Human herpesvirus 2 |
| Molecular weight | Theoretical: 50.542617 KDa |
| Sequence | String: MKTKPLPTAP MAWAESAVET TTSPRELAGH APLRRVLRPP IARRDGPVLL GDRAPRRTAS TMWLLGIDPA ESSPGTRATR DDTEQAVDK ILRGARRAGG LTVPGAPRYH LTRQVTLTDL CQPNAERAGA LLLALRHPTD LPHLARHRAP PGRQTERLAE A WGQLLEAS ...String: MKTKPLPTAP MAWAESAVET TTSPRELAGH APLRRVLRPP IARRDGPVLL GDRAPRRTAS TMWLLGIDPA ESSPGTRATR DDTEQAVDK ILRGARRAGG LTVPGAPRYH LTRQVTLTDL CQPNAERAGA LLLALRHPTD LPHLARHRAP PGRQTERLAE A WGQLLEAS ALGSGRAESG CARAGLVSFN FLVAACTAAY DARDAAEAVR AHITTNYGGT RAGARLDRFS ECLRAMVHTH VF PHEVMRF FGGLVSWVTQ DELASVTAVC SGPQEATHTG HPGRPCSAVT IPACAFVDLD AELCLGGPGA AFLYLVFTYR QCR DQELCC VYVVKSQLPP RGLEAALERL FGRLRITNTI HGAEDMTPPP PNRNVDFPLA VLAASSQSPR CSASQVTNPQ FVDR LYRWQ PDLRGRPTAR TCTYAAFAEL GVMPDDSPRC LHRTERFGAV GVPVVILEGV VWRPGGWRAC A UniProtKB: Capsid triplex subunit 1 |
-Macromolecule #4: Small capsomere-interacting protein
| Macromolecule | Name: Small capsomere-interacting protein / type: protein_or_peptide / ID: 4 / Number of copies: 15 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human herpesvirus 2 Human herpesvirus 2 |
| Molecular weight | Theoretical: 12.147707 KDa |
| Sequence | String: MAAPQFHRPS TITADNVRAL GMRGLVLATN NAQFIMDNSY PHPHGTQGAV REFLRGQAAA LTDLGVTHAN NTFAPQPMFA GDAAAEWLR PSFGLKRTYS PFVVRDPKTP STP UniProtKB: Small capsomere-interacting protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 1.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 45000 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: COMMON LINE |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)