+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | ATP-bound ADP-Glucose Pyrophosphorylase | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | heterotetramer / ADPG / PLANT PROTEIN / amylosynthesis | |||||||||
| Function / homology |  Function and homology information Function and homology informationheterotetrameric ADPG pyrophosphorylase complex / glucose-1-phosphate adenylyltransferase complex / glucose-1-phosphate adenylyltransferase / glucose-1-phosphate adenylyltransferase activity / starch biosynthetic process / photoperiodism, flowering / apoplast / chloroplast envelope / glycogen biosynthetic process / chloroplast stroma ...heterotetrameric ADPG pyrophosphorylase complex / glucose-1-phosphate adenylyltransferase complex / glucose-1-phosphate adenylyltransferase / glucose-1-phosphate adenylyltransferase activity / starch biosynthetic process / photoperiodism, flowering / apoplast / chloroplast envelope / glycogen biosynthetic process / chloroplast stroma / plastid / chloroplast / ATP binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.81 Å | |||||||||
 Authors Authors | Wu YT / Lin HJ / Fan MR | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: ATP-bound ADP-Glucose Pyrophosphorylase Authors: Wu YT / Lin HJ / Fan MR | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_67276.map.gz emd_67276.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-67276-v30.xml emd-67276-v30.xml emd-67276.xml emd-67276.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_67276_fsc.xml emd_67276_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_67276.png emd_67276.png | 25.4 KB | ||
| Masks |  emd_67276_msk_1.map emd_67276_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-67276.cif.gz emd-67276.cif.gz | 6.1 KB | ||
| Others |  emd_67276_half_map_1.map.gz emd_67276_half_map_1.map.gz emd_67276_half_map_2.map.gz emd_67276_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-67276 http://ftp.pdbj.org/pub/emdb/structures/EMD-67276 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-67276 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-67276 | HTTPS FTP |
-Related structure data
| Related structure data |  9xusMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_67276.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_67276.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.055 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_67276_msk_1.map emd_67276_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_67276_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_67276_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ATP-bound ADP-Glucose Pyrophosphorylase
| Entire | Name: ATP-bound ADP-Glucose Pyrophosphorylase |
|---|---|
| Components |
|
-Supramolecule #1: ATP-bound ADP-Glucose Pyrophosphorylase
| Supramolecule | Name: ATP-bound ADP-Glucose Pyrophosphorylase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Glucose-1-phosphate adenylyltransferase small subunit, chloroplastic
| Macromolecule | Name: Glucose-1-phosphate adenylyltransferase small subunit, chloroplastic type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: glucose-1-phosphate adenylyltransferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 51.517086 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSHHHHHHH HGSGSGSGSG SVSDSQNSQT CLDPDASSSV LGIILGGGAG TRLYPLTKKR AKPAVPLGAN YRLIDIPVSN CLNSNISKI YVLTQFNSAS LNRHLSRAYA SNMGGYKNEG FVEVLAAQQS PENPNWFQGT ADAVRQYLWL FEEHNVLEYL I LAGDHLYR ...String: MGSHHHHHHH HGSGSGSGSG SVSDSQNSQT CLDPDASSSV LGIILGGGAG TRLYPLTKKR AKPAVPLGAN YRLIDIPVSN CLNSNISKI YVLTQFNSAS LNRHLSRAYA SNMGGYKNEG FVEVLAAQQS PENPNWFQGT ADAVRQYLWL FEEHNVLEYL I LAGDHLYR MDYEKFIQAH RETDADITVA ALPMDEQRAT AFGLMKIDEE GRIIEFAEKP KGEHLKAMKV DTTILGLDDQ RA KEMPFIA SMGIYVVSRD VMLDLLRNQF PGANDFGSEV IPGATSLGLR VQAYLYDGYW EDIGTIEAFY NANLGITKKP VPD FSFYDR SAPIYTQPRY LPPSKMLDAD VTDSVIGEGC VIKNCKIHHS VVGLRSCISE GAIIEDSLLM GADYYETATE KSLL SAKGS VPIGIGKNSH IKRAIIDKNA RIGDNVKIIN SDNVQEAARE TDGYFIKSGI VTVIKDALIP TGTVI UniProtKB: Glucose-1-phosphate adenylyltransferase small subunit, chloroplastic |
-Macromolecule #2: Glucose-1-phosphate adenylyltransferase large subunit 1, chloroplastic
| Macromolecule | Name: Glucose-1-phosphate adenylyltransferase large subunit 1, chloroplastic type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: glucose-1-phosphate adenylyltransferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 52.535211 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSWSHPQFE KGSGSGSGSG SSLNSVAGES KVQELETEKR DPRTVASIIL GGGAGTRLFP LTKRRAKPAV PIGGAYRLID VPMSNCINS GINKVYILTQ YNSASLNRHL ARAYNSNGLG FGDGYVEVLA ATQTPGESGK RWFQGTADAV RQFHWLFEDA R SKDIEDVL ...String: MGSWSHPQFE KGSGSGSGSG SSLNSVAGES KVQELETEKR DPRTVASIIL GGGAGTRLFP LTKRRAKPAV PIGGAYRLID VPMSNCINS GINKVYILTQ YNSASLNRHL ARAYNSNGLG FGDGYVEVLA ATQTPGESGK RWFQGTADAV RQFHWLFEDA R SKDIEDVL ILSGDHLYRM DYMDFIQDHR QSGADISISC IPIDDRRASD FGLMKIDDKG RVISFSEKPK GDDLKAMAVD TT ILGLSKE EAEKKPYIAS MGVYVFKKEI LLNLLRWRFP TANDFGSEII PFSAKEFYVN AYLFNDYWED IGTIRSFFEA NLA LTEHPG AFSFYDAAKP IYTSRRNLPP SKIDNSKLID SIISHGSFLT NCLIEHSIVG IRSRVGSNVQ LKDTVMLGAD YYET EAEVA ALLAEGNVPI GIGENTKIQE CIIDKNARVG KNVIIANSEG IQEADRSSDG FYIRSGITVI LKNSVIKDGV VI UniProtKB: Glucose-1-phosphate adenylyltransferase large subunit 1, chloroplastic |
-Macromolecule #3: 3-PHOSPHOGLYCERIC ACID
| Macromolecule | Name: 3-PHOSPHOGLYCERIC ACID / type: ligand / ID: 3 / Number of copies: 4 / Formula: 3PG |
|---|---|
| Molecular weight | Theoretical: 186.057 Da |
| Chemical component information | 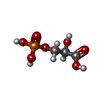 ChemComp-3PG: |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 8 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































