+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6595 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM of the C. elegans Ndc80 complex bound to microtubules | |||||||||
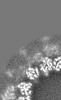
 Map data Map data | Helical reconstruction of the C. elegans Ndc80 complex bound to microtubules in the presence of the SKA1 complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Kinetochore protein complex | |||||||||
| Biological species | unidentified (others) /  | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 4.06 Å | |||||||||
 Authors Authors | Wilson-Kubalek EM / Cheeseman IM / Milligan RA | |||||||||
 Citation Citation |  Journal: Mol Biol Cell / Year: 2016 Journal: Mol Biol Cell / Year: 2016Title: Structural comparison of the Caenorhabditis elegans and human Ndc80 complexes bound to microtubules reveals distinct binding behavior. Authors: Elizabeth M Wilson-Kubalek / Iain M Cheeseman / Ronald A Milligan /  Abstract: During cell division, kinetochores must remain tethered to the plus ends of dynamic microtubule polymers. However, the molecular basis for robust kinetochore-microtubule interactions remains poorly ...During cell division, kinetochores must remain tethered to the plus ends of dynamic microtubule polymers. However, the molecular basis for robust kinetochore-microtubule interactions remains poorly understood. The conserved four-subunit Ndc80 complex plays an essential and direct role in generating dynamic kinetochore-microtubule attachments. Here we compare the binding of theCaenorhabditis elegansand human Ndc80 complexes to microtubules at high resolution using cryo-electron microscopy reconstructions. Despite the conserved roles of the Ndc80 complex in diverse organisms, we find that the attachment mode of these complexes for microtubules is distinct. The human Ndc80 complex binds every tubulin monomer along the microtubule protofilament, whereas theC. elegansNdc80 complex binds more tightly to β-tubulin. In addition, theC. elegansNdc80 complex tilts more toward the adjacent protofilament. These structural differences in the Ndc80 complex between different species may play significant roles in the nature of kinetochore-microtubule interactions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6595.map.gz emd_6595.map.gz | 15.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6595-v30.xml emd-6595-v30.xml emd-6595.xml emd-6595.xml | 9.6 KB 9.6 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6595.gif 400_6595.gif 80_6595.gif 80_6595.gif | 136.2 KB 8.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6595 http://ftp.pdbj.org/pub/emdb/structures/EMD-6595 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6595 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6595 | HTTPS FTP |
-Validation report
| Summary document |  emd_6595_validation.pdf.gz emd_6595_validation.pdf.gz | 78.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6595_full_validation.pdf.gz emd_6595_full_validation.pdf.gz | 77.5 KB | Display | |
| Data in XML |  emd_6595_validation.xml.gz emd_6595_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6595 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6595 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6595 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6595 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6595.map.gz / Format: CCP4 / Size: 16.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6595.map.gz / Format: CCP4 / Size: 16.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Helical reconstruction of the C. elegans Ndc80 complex bound to microtubules in the presence of the SKA1 complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
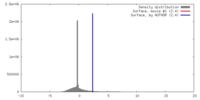
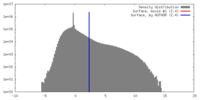
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : full-length C. elegans Ndc80 complex bound to microtubules
| Entire | Name: full-length C. elegans Ndc80 complex bound to microtubules |
|---|---|
| Components |
|
-Supramolecule #1000: full-length C. elegans Ndc80 complex bound to microtubules
| Supramolecule | Name: full-length C. elegans Ndc80 complex bound to microtubules type: sample / ID: 1000 / Oligomeric state: helical / Number unique components: 2 |
|---|
-Supramolecule #1: microtubule
| Supramolecule | Name: microtubule / type: organelle_or_cellular_component / ID: 1 / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism: unidentified (others) |
-Macromolecule #1: Ndc80
| Macromolecule | Name: Ndc80 / type: protein_or_peptide / ID: 1 Details: C. elegans Ndc80 complex was bound to the microtubule in the presence of the SKA1 complex Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 / Details: 80 mM PIPES, pH 6.8, 1 mM MgCl2, 1 mM EGTA |
|---|---|
| Grid | Details: 400-mesh C-flat grids (Protochips, Inc) containing 2.0 micron holes separated by 2.0 micron spacing |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 120 K / Instrument: HOMEMADE PLUNGER / Method: Blotted back of grid for 3 seconds. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Date | Sep 23, 2015 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Digitization - Sampling interval: 1.31 µm / Number real images: 984 / Average electron dose: 40 e/Å2 / Details: Each image is an average of 40 frames. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | EMAN2 with IHRSR adapted for microtubules with a dimer repeat and FREALIGN was used. |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 11.092 Å Applied symmetry - Helical parameters - Δ&Phi: 23.825 ° Applied symmetry - Helical parameters - Axial symmetry: C15 (15 fold cyclic) Resolution.type: BY AUTHOR / Resolution: 4.06 Å / Resolution method: OTHER / Software - Name: EMAN2, FREALIGN, IHRSR Details: The final map was calculated from two averaged data sets. |
| CTF correction | Details: CTFIND v3 |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)