+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PvdL-E2-C3-A3-PCP3 in complex with MLP (NRPS cross-module) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Nonribosomal peptide synthetase / LIGASE / BIOSYNTHETIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpyoverdine biosynthetic process / monocarboxylic acid biosynthetic process / Actinobacterium-type cell wall biogenesis / siderophore biosynthetic process / amino acid activation for nonribosomal peptide biosynthetic process / lipid biosynthetic process / ligase activity / catalytic activity / phosphopantetheine binding / antibiotic biosynthetic process ...pyoverdine biosynthetic process / monocarboxylic acid biosynthetic process / Actinobacterium-type cell wall biogenesis / siderophore biosynthetic process / amino acid activation for nonribosomal peptide biosynthetic process / lipid biosynthetic process / ligase activity / catalytic activity / phosphopantetheine binding / antibiotic biosynthetic process / fatty acid metabolic process / transferase activity / cytosol Similarity search - Function | |||||||||
| Biological species |  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.33 Å | |||||||||
 Authors Authors | Cao W / Wang J / Wang Z | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Synth / Year: 2026 Journal: Nat Synth / Year: 2026Title: Analysis of heterocycle formation and stereochemical control by a non-ribosomal peptide synthetase condensation domain Authors: Cao W / Wang J / Chen SL / Li D / Wang X / Deng Z / Liang J / Wang Z | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_64387.map.gz emd_64387.map.gz | 95.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-64387-v30.xml emd-64387-v30.xml emd-64387.xml emd-64387.xml | 22.6 KB 22.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_64387_fsc.xml emd_64387_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_64387.png emd_64387.png | 71.1 KB | ||
| Filedesc metadata |  emd-64387.cif.gz emd-64387.cif.gz | 6.8 KB | ||
| Others |  emd_64387_additional_1.map.gz emd_64387_additional_1.map.gz emd_64387_additional_2.map.gz emd_64387_additional_2.map.gz emd_64387_half_map_1.map.gz emd_64387_half_map_1.map.gz emd_64387_half_map_2.map.gz emd_64387_half_map_2.map.gz | 96.3 MB 6.1 MB 81.2 MB 81.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-64387 http://ftp.pdbj.org/pub/emdb/structures/EMD-64387 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64387 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64387 | HTTPS FTP |
-Related structure data
| Related structure data |  9up2MC  9lcoC  9lcpC  9lcqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_64387.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_64387.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #2
| File | emd_64387_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_64387_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_64387_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_64387_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PvdL-E2-C3-A3-PCP3 in complex with MLP (NRPS cross-module)
| Entire | Name: PvdL-E2-C3-A3-PCP3 in complex with MLP (NRPS cross-module) |
|---|---|
| Components |
|
-Supramolecule #1: PvdL-E2-C3-A3-PCP3 in complex with MLP (NRPS cross-module)
| Supramolecule | Name: PvdL-E2-C3-A3-PCP3 in complex with MLP (NRPS cross-module) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
| Molecular weight | Theoretical: 186 kDa/nm |
-Macromolecule #1: PvdL
| Macromolecule | Name: PvdL / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
| Molecular weight | Theoretical: 172.528516 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DSALTPIQHW FFDLPLARRE HWNQALLLQP RQAIDLGLLR KSLQRLVEQH DALRLAFRQV DGEWLAQHRP LREQELLWHV PVQSFDECA ELFAKAQRSL DLEQGPLLRA VLVDGPAGEQ RLLLAIHHLV VDGVSWRVLL EDLQQVYRQF AEGAEPALPA K TSAFRDWA ...String: DSALTPIQHW FFDLPLARRE HWNQALLLQP RQAIDLGLLR KSLQRLVEQH DALRLAFRQV DGEWLAQHRP LREQELLWHV PVQSFDECA ELFAKAQRSL DLEQGPLLRA VLVDGPAGEQ RLLLAIHHLV VDGVSWRVLL EDLQQVYRQF AEGAEPALPA K TSAFRDWA GRLQAYAGSE SLREELGWWQ ARLGGQPVEW PCDRPQGDNR EALAESVSLR LDPQRTRQLL QQAPAAYRTQ VN DLLLTAL ARVLCRWSGQ PSTLVQLEGH GREALFDDID LTRSVGWFTS AYPLRLTPAQ SPGESIKAIK EQLRAVPHKG LGY GVLRYL ADPAVRQAMA ALPTAPITFN YLGQFDQSFA DALFQPLDQP TGPIHDEQAP LPNELSVDGQ VYGGELVLRW TYSR ERYDA RTVNELAQAY LAELQALIEH CLEDGAGGLT PSDFPLAQLS QAQLDALAVP AGEIEDVYPL TPMQEGLLLH TLLEP GTGI YYMQDRYRID SPLDPERFAA AWQAVVARHE ALRASFVWNA GETMLQVIHK PGRTRIEFLD WSELPEDGHE ERLQAL HKR EREAGFDLLE QPPFHLRLIR LGEARYWFMM SNHHILIDAW CRGLLMNDFF EIYSALGESR PANLPTPPRY RDYIAWL QR QDLEQSRRWW SESLRGFERP TLVPSDRPFL REHAGESGGM IVGDRYTRLD AADGARLREL AQRYQLTVNT FAQAAWAL T LRRFSGERDV LFGVTVAGRP VGMPEMQRTV GLFINSIPLR VQMPAAGQRC TVREWLNRLF ERNLELREHE HLPLVAIQE SSELPKGQPL FDSLFVFENA PVEVSVLDRA QSLNASSDSG RTHTNFPLTV VCYPGDDLGL HLSYDQRYFE APTVERLLGE FKRLLLALA DGFHGELEAL PLLGEDERDF LLDGCNRSAR DYPLEQGYVR LFEAQVAAHP QRIAASCLEQ RWSYAELNRR A NRLGHALR AAGVGIDQPV ALLAERGLDL LGMIVGSFKA GAGYLPLDPG HPTQRLTRIV ELSRTLVLVC TQACREQALA LF DELGCVD RPRLLVWDEI QQGEGAEHDP QVYSGPQNLA YVIYTSGSTG LPKGVMVEQA GMLNNQLSKV PYLELDENDV IAQ TASQSF DISVWQFLAA PLFGARVAIV PNAVAHDPQG LLAHVGEQGI TVLESVPSLI QGMLAEERQA LDGLRWMLPT GEAM PPELA RQWLKRYPRI GLVNAYGPAE CSDDVAFFRV DLASTESTYL PIGSPTDNNR LYLLGAGADD AFELVPLGAV GELCV AGTG VGRGYVGDPL RTAQAFVPHP FGAPGERLYR TGDLARRRAD GVLEYVGRID HQVKIRGFRI ELGEIEARLH ERADVR EAA VAVQEGANGK YLVGYLVPGE TPRSSADSPA GLMVEQGAWF ERIKQQLRAD LPDYMVPLHW LVLDRMPLNA NGKLDRK AL PALDIGQMQN QAYQAPRNEL EETLARIWAE VLKVERVGVF DNFFELGGH(4HH) LLATQIASRV QKALQRNVPL RAMF ECTTV EELASYIESL APSEI UniProtKB: PvdL |
-Macromolecule #2: MbtH-like domain-containing protein
| Macromolecule | Name: MbtH-like domain-containing protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
| Molecular weight | Theoretical: 7.28819 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DDIQFQVVVN HEEQYSIWPE YKEIPQGWRA AGKSGLKKDC LAYIEEVWTD MRPLSLRQHM D UniProtKB: MbtH-like domain-containing protein |
-Macromolecule #3: 2,4-DIAMINOBUTYRIC ACID
| Macromolecule | Name: 2,4-DIAMINOBUTYRIC ACID / type: ligand / ID: 3 / Number of copies: 2 / Formula: DAB |
|---|---|
| Molecular weight | Theoretical: 118.134 Da |
| Chemical component information | 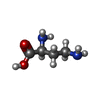 ChemComp-DAB: |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 1 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 64000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















































