+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CTP synthase DON-state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CTP synthase / CTPS / LIGASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationInterconversion of nucleotide di- and triphosphates / larval lymph gland hemopoiesis / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / CTP biosynthetic process / ATP binding / identical protein binding ...Interconversion of nucleotide di- and triphosphates / larval lymph gland hemopoiesis / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / CTP biosynthetic process / ATP binding / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.99 Å | |||||||||
 Authors Authors | Guo CJ | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: CTP synthase DON-state Authors: Guo CJ | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_63990.map.gz emd_63990.map.gz | 8.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-63990-v30.xml emd-63990-v30.xml emd-63990.xml emd-63990.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_63990_fsc.xml emd_63990_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_63990.png emd_63990.png | 112.1 KB | ||
| Masks |  emd_63990_msk_1.map emd_63990_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-63990.cif.gz emd-63990.cif.gz | 5.8 KB | ||
| Others |  emd_63990_half_map_1.map.gz emd_63990_half_map_1.map.gz emd_63990_half_map_2.map.gz emd_63990_half_map_2.map.gz | 200.4 MB 200.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-63990 http://ftp.pdbj.org/pub/emdb/structures/EMD-63990 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63990 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63990 | HTTPS FTP |
-Related structure data
| Related structure data |  9uarMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_63990.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_63990.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.707 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_63990_msk_1.map emd_63990_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_63990_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_63990_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : DON-state CTP synthase
| Entire | Name: DON-state CTP synthase |
|---|---|
| Components |
|
-Supramolecule #1: DON-state CTP synthase
| Supramolecule | Name: DON-state CTP synthase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: CTP synthase
| Macromolecule | Name: CTP synthase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: CTP synthase (glutamine hydrolysing) |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 62.443656 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKYILVTGGV ISGVGKGVIA SSFGTLLKSC GLDVTSIKID PYINIDAGTF SPYEHGEVYV LDDGAEVDLD LGNYERFLDV TLHRDNNIT TGKIYKLVIE KERTGEYLGK TVQVVPHITD AIQEWVERVA QTPVQGSSKP QVCIVELGGT IGDIEGMPFV E AFRQFQFR ...String: MKYILVTGGV ISGVGKGVIA SSFGTLLKSC GLDVTSIKID PYINIDAGTF SPYEHGEVYV LDDGAEVDLD LGNYERFLDV TLHRDNNIT TGKIYKLVIE KERTGEYLGK TVQVVPHITD AIQEWVERVA QTPVQGSSKP QVCIVELGGT IGDIEGMPFV E AFRQFQFR VKRENFCLAH VSLVPLPKAT GEPKTKPTQS SVRELRGCGL SPDLIVCRSE KPIGLEVKEK ISNFCHVGPD QV ICIHDLN SIYHVPLLME QNGVIEYLNE RLQLNIDMSK RTKCLQQWRD LARRTETVRR EVCIAVVGKY TKFTDSYASV VKA LQHAAL AVNRKLELVF IESCLLEEET LHSEPSKYHK EWQKLCDSHG ILVPGGFGSR GMEGKIRACQ WARENQKPLL GICL GLQAA VIEFARNKLG LKDANTTEID PNTANALVID MPEHHTGQLG GTMRLGKRIT VFSDGPSVIR QLYGNPKSVQ ERHRH RYEV NPKYVHLLEE QGMRFVGTDV DKTRMEIIEL SGHPYFVATQ YHPEYLSRPL KPSPPFLGLI LASVDRLNQY IQ UniProtKB: CTP synthase |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 4 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #3: [[(2~{R},3~{S},4~{R},5~{R})-3,4-bis(oxidanyl)-5-(2-oxidanyl-4-pho...
| Macromolecule | Name: [[(2~{R},3~{S},4~{R},5~{R})-3,4-bis(oxidanyl)-5-(2-oxidanyl-4-phosphonooxy-pyrimidin-1-yl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] phosphono hydrogen phosphate type: ligand / ID: 3 / Number of copies: 4 / Formula: 5ZL |
|---|---|
| Molecular weight | Theoretical: 565.129 Da |
| Chemical component information | 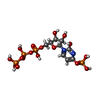 ChemComp-5ZL: |
-Macromolecule #4: GUANOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 4 / Formula: GTP |
|---|---|
| Molecular weight | Theoretical: 523.18 Da |
| Chemical component information |  ChemComp-GTP: |
-Macromolecule #5: 5-OXO-L-NORLEUCINE
| Macromolecule | Name: 5-OXO-L-NORLEUCINE / type: ligand / ID: 5 / Number of copies: 4 / Formula: ONL |
|---|---|
| Molecular weight | Theoretical: 145.156 Da |
| Chemical component information | 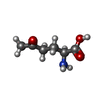 ChemComp-ONL: |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 16 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #7: water
| Macromolecule | Name: water / type: ligand / ID: 7 / Number of copies: 888 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)














































