[English] 日本語
 Yorodumi
Yorodumi- EMDB-63941: Tetrameric cystathionine beta-synthase of Mycobacterium tuberculo... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tetrameric cystathionine beta-synthase of Mycobacterium tuberculosis bound to AOAA | |||||||||
 Map data Map data | AOAA-bound MtbCBS map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Inhibitor / Lyase / Complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationcystathionine beta-synthase / cystathionine beta-synthase activity / : / cysteine synthase activity / : / peptidoglycan-based cell wall / extracellular region / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) (bacteria) Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.96 Å | |||||||||
 Authors Authors | Roy A / Polepalli S / Mondal B / Dutta S | |||||||||
| Funding support |  India, 2 items India, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Tetrameric cystathionine beta-synthase of Mycobacterium tuberculosis bound to AOAA Authors: Roy A / Polepalli S / Mondal B / Dutta S | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_63941.map.gz emd_63941.map.gz | 89.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-63941-v30.xml emd-63941-v30.xml emd-63941.xml emd-63941.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_63941.png emd_63941.png | 30.1 KB | ||
| Filedesc metadata |  emd-63941.cif.gz emd-63941.cif.gz | 5.7 KB | ||
| Others |  emd_63941_half_map_1.map.gz emd_63941_half_map_1.map.gz emd_63941_half_map_2.map.gz emd_63941_half_map_2.map.gz | 88 MB 88 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-63941 http://ftp.pdbj.org/pub/emdb/structures/EMD-63941 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63941 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63941 | HTTPS FTP |
-Related structure data
| Related structure data |  9u7oMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_63941.map.gz / Format: CCP4 / Size: 95 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_63941.map.gz / Format: CCP4 / Size: 95 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AOAA-bound MtbCBS map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.92 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: AOAA-bound MtbCBS half-map A
| File | emd_63941_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AOAA-bound MtbCBS half-map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: AOAA-bound MtbCBS half-map B
| File | emd_63941_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AOAA-bound MtbCBS half-map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The ternary complex of AOAA-bound to MtbCBS
| Entire | Name: The ternary complex of AOAA-bound to MtbCBS |
|---|---|
| Components |
|
-Supramolecule #1: The ternary complex of AOAA-bound to MtbCBS
| Supramolecule | Name: The ternary complex of AOAA-bound to MtbCBS / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) (bacteria) Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) (bacteria) |
-Macromolecule #1: Probable cystathionine beta-synthase Rv1077
| Macromolecule | Name: Probable cystathionine beta-synthase Rv1077 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: cystathionine beta-synthase |
|---|---|
| Source (natural) | Organism:  Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) (bacteria) Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) (bacteria) |
| Molecular weight | Theoretical: 50.284848 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MARIAQHISE LIGGTPLVRL NSVVPDGAGT VAAKVEYLNP GGSSKDRIAV KMIEAAEASG QLKPGGTIVE PTSGNTGVGL ALVAQRRGY KCVFVCPDKV SEDKRNVLIA YGAEVVVCPT AVPPHDPASY YSVSDRLVRD IDGAWKPDQY ANPEGPASHY V TTGPEIWA ...String: MARIAQHISE LIGGTPLVRL NSVVPDGAGT VAAKVEYLNP GGSSKDRIAV KMIEAAEASG QLKPGGTIVE PTSGNTGVGL ALVAQRRGY KCVFVCPDKV SEDKRNVLIA YGAEVVVCPT AVPPHDPASY YSVSDRLVRD IDGAWKPDQY ANPEGPASHY V TTGPEIWA DTEGKVTHFV AGIGTGGTIT GAGRYLKEVS GGRVRIVGAD PEGSVYSGGA GRPYLVEGVG EDFWPAAYDP SV PDEIIAV SDSDSFDMTR RLAREEAMLV GGSCGMAVVA ALKVAEEAGP DALIVVLLPD GGRGYMSKIF NDAWMSSYGF LRS RLDGST EQSTVGDVLR RKSGALPALV HTHPSETVRD AIGILREYGV SQMPVVGAEP PVMAGEVAGS VSERELLSAV FEGR AKLAD AVSAHMSPPL RMIGAGELVS AAGKALRDWD ALMVVEEGKP VGVITRYDLL GFLSEGAGRR KLAAALEHHH HHH UniProtKB: Probable cystathionine beta-synthase Rv1077 |
-Macromolecule #2: 4'-DEOXY-4'-ACETYLYAMINO-PYRIDOXAL-5'-PHOSPHATE
| Macromolecule | Name: 4'-DEOXY-4'-ACETYLYAMINO-PYRIDOXAL-5'-PHOSPHATE / type: ligand / ID: 2 / Number of copies: 4 / Formula: IK2 |
|---|---|
| Molecular weight | Theoretical: 322.208 Da |
| Chemical component information | 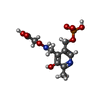 ChemComp-IK2: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































