+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dimer structure of a glycosyltransferase | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | glycosyltransferase / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationprocollagen galactosyltransferase / procollagen galactosyltransferase activity / positive regulation of collagen fibril organization / Collagen biosynthesis and modifying enzymes / protein O-linked glycosylation / collagen fibril organization / endoplasmic reticulum lumen / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Yu HJ / Zhang M / Sun HH / Liu XT | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Dimer structure of a glycosyltransferase Authors: Yu HJ / Zhang M / Sun HH / Liu XT | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_63937.map.gz emd_63937.map.gz | 49.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-63937-v30.xml emd-63937-v30.xml emd-63937.xml emd-63937.xml | 16 KB 16 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_63937_fsc.xml emd_63937_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_63937.png emd_63937.png | 84.8 KB | ||
| Filedesc metadata |  emd-63937.cif.gz emd-63937.cif.gz | 5.7 KB | ||
| Others |  emd_63937_half_map_1.map.gz emd_63937_half_map_1.map.gz emd_63937_half_map_2.map.gz emd_63937_half_map_2.map.gz | 40.7 MB 40.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-63937 http://ftp.pdbj.org/pub/emdb/structures/EMD-63937 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63937 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63937 | HTTPS FTP |
-Related structure data
| Related structure data |  9u7iMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_63937.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_63937.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.076 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_63937_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_63937_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : glycosyltransferase
| Entire | Name: glycosyltransferase |
|---|---|
| Components |
|
-Supramolecule #1: glycosyltransferase
| Supramolecule | Name: glycosyltransferase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Procollagen galactosyltransferase 1
| Macromolecule | Name: Procollagen galactosyltransferase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: procollagen galactosyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 71.069969 KDa |
| Recombinant expression | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
| Sequence | String: APPGADAYFP EERWSPESPL QAPRVLIALL ARNAAHALPT TLGALERLRH PRERTALWVA TDHNMDNTST VLREWLVAVK SLYHSVEWR PAEEPRSYPD EEGPKHWSDS RYEHVMKLRQ AALKSARDMW ADYILFVDAD NLILNPDTLS LLIAENKTVV A PMLDSRAA ...String: APPGADAYFP EERWSPESPL QAPRVLIALL ARNAAHALPT TLGALERLRH PRERTALWVA TDHNMDNTST VLREWLVAVK SLYHSVEWR PAEEPRSYPD EEGPKHWSDS RYEHVMKLRQ AALKSARDMW ADYILFVDAD NLILNPDTLS LLIAENKTVV A PMLDSRAA YSNFWCGMTS QGYYKRTPAY IPIRKRDRRG CFAVPMVHST FLIDLRKAAS RNLAFYPPHP DYTWSFDDII VF AFSCKQA EVQMYVCNKE EYGFLPVPLR AHSTLQDEAE SFMHVQLEVM VKHPPAEPSR FISAPTKTPD KMGFDEVFMI NLR RRQDRR ERMLRALQAQ EIECRLVEAV DGKAMNTSQV EALGIQMLPG YRDPYHGRPL TKGELGCFLS HYNIWKEVVD RGLQ KSLVF EDDLRFEIFF KRRLMNLMRD VEREGLDWDL IYVGRKRMQV EHPEKAVPRV RNLVEADYSY WTLAYVISLQ GARKL LAAE PLSKMLPVDE FLPVMFDKHP VSEYKAHFSL RNLHAFSVEP LLIYPTHYTG DDGYVSDTET SVVWNNEHVK TDWDRA KSQ KMREQQALSR EAKNSDVLQS PLDSAARGTL EVLFQGPLGS HHHHHHHHHH UniProtKB: Procollagen galactosyltransferase 1 |
-Macromolecule #2: GALACTOSE-URIDINE-5'-DIPHOSPHATE
| Macromolecule | Name: GALACTOSE-URIDINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 2 / Formula: GDU |
|---|---|
| Molecular weight | Theoretical: 566.302 Da |
| Chemical component information | 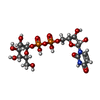 ChemComp-GDU: |
-Macromolecule #3: MANGANESE (II) ION
| Macromolecule | Name: MANGANESE (II) ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: MN |
|---|---|
| Molecular weight | Theoretical: 54.938 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.1 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





































