[English] 日本語
 Yorodumi
Yorodumi- EMDB-63807: Plant chloroplast dicarboxylate transporter AtDiT2.1 bound with malate -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Plant chloroplast dicarboxylate transporter AtDiT2.1 bound with malate | |||||||||
 Map data Map data | main_map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | chloroplast / dicarboxylate transporters / C/N coupling / C4 photosynthesis / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationoxaloacetate transmembrane transporter activity / oxaloacetate transport / malate transmembrane transport / malate transmembrane transporter activity / ammonia assimilation cycle / chloroplast inner membrane / L-glutamate transmembrane transporter activity / L-glutamate transmembrane transport / chloroplast thylakoid / chloroplast envelope ...oxaloacetate transmembrane transporter activity / oxaloacetate transport / malate transmembrane transport / malate transmembrane transporter activity / ammonia assimilation cycle / chloroplast inner membrane / L-glutamate transmembrane transporter activity / L-glutamate transmembrane transport / chloroplast thylakoid / chloroplast envelope / response to nematode / plastid / chloroplast Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.72 Å | |||||||||
 Authors Authors | Yang Z / Zhang P | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Plant Cell / Year: 2026 Journal: Plant Cell / Year: 2026Title: Substrate specificity and transport mechanism of the chloroplast dicarboxylate transporters DiT1 and DiT2. Authors: Zhao Yang / Xue Zhang / Jingtao Zheng / Shunhao Zhou / Ming-Ju Amy Lyu / Miaolian Ma / Xin-Guang Zhu / Fang Yu / Peng Zhang /  Abstract: Dicarboxylate transporters (DiTs) mediate the exchange of dicarboxylates across the chloroplast inner membrane, playing critical roles in C/N coupling, photorespiration, chloroplast redox ...Dicarboxylate transporters (DiTs) mediate the exchange of dicarboxylates across the chloroplast inner membrane, playing critical roles in C/N coupling, photorespiration, chloroplast redox homeostasis, and C4 photosynthesis. DiT1 and DiT2 are Na⁺-independent exchangers of the solute carrier 13 (SLC13) family, and exhibit overlapping yet distinct substrate specificities: DiT1 transports 2-oxoglutarate, malate, and oxaloacetate, while DiT2 additionally transports glutamate and aspartate. However, the structural determinants of their substrate specificity and transport mechanism remain unclear. Here we determined cryo-electron microscopy structures of Arabidopsis thaliana DiT1 and DiT2.1 bound to diverse substrates in dual conformational states. Structural analyses revealed that AtDiT1 possesses a singular dicarboxylate-binding site that is electrostatically incompatible with amino acid substrates, whereas AtDiT2.1 has two distinct sites to accommodate C4- and C5-dicarboxylates, thus allowing amino acids to bind without electrostatic repulsion. Phylogenetic analysis identified an A226S substitution in the substrate-binding site of DiT1, emerging during evolution in the charophyte ancestor of land plants. This substitution enhances oxaloacetate binding affinity in DiT1, which may have improved adaptation to terrestrial environments. Additionally, two conserved positively charged residues in DiTs functionally mimic Na⁺ used by SLC13 co-transporters, thereby enabling a Na⁺-independent elevator-type transport mechanism. These findings provide critical structural and mechanistic insights into the functional divergence of plant DiTs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_63807.map.gz emd_63807.map.gz | 78.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-63807-v30.xml emd-63807-v30.xml emd-63807.xml emd-63807.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_63807_fsc.xml emd_63807_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_63807.png emd_63807.png | 87.1 KB | ||
| Filedesc metadata |  emd-63807.cif.gz emd-63807.cif.gz | 5.9 KB | ||
| Others |  emd_63807_half_map_1.map.gz emd_63807_half_map_1.map.gz emd_63807_half_map_2.map.gz emd_63807_half_map_2.map.gz | 77.4 MB 77.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-63807 http://ftp.pdbj.org/pub/emdb/structures/EMD-63807 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63807 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63807 | HTTPS FTP |
-Related structure data
| Related structure data |  9mcvMC  9mcrC  9mcsC  9mctC  9mcuC  9u32C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_63807.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_63807.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main_map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.932 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half B
| File | emd_63807_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half_B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half A
| File | emd_63807_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half_A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Plant chloroplast dicarboxylate transporter AtDiT2.1 bound with malate
| Entire | Name: Plant chloroplast dicarboxylate transporter AtDiT2.1 bound with malate |
|---|---|
| Components |
|
-Supramolecule #1: Plant chloroplast dicarboxylate transporter AtDiT2.1 bound with malate
| Supramolecule | Name: Plant chloroplast dicarboxylate transporter AtDiT2.1 bound with malate type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Dicarboxylate transporter 2.1, chloroplastic
| Macromolecule | Name: Dicarboxylate transporter 2.1, chloroplastic / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 55.765598 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYKDDDDKD YKDDDDKDYK DDDDKLEIAA APQDNAPPPP PPSPSPSPSP QGAKLIPLIL SISVGLILRF AVPVPEGVTP QGWQLLSIF LSTIAGLVLS PLPVGAWAFI GLTASIVTKT LSFSAAFSAF TSEVIWLIVI SFFFARGFVK TGLGDRIATY F VKWLGKST ...String: MDYKDDDDKD YKDDDDKDYK DDDDKLEIAA APQDNAPPPP PPSPSPSPSP QGAKLIPLIL SISVGLILRF AVPVPEGVTP QGWQLLSIF LSTIAGLVLS PLPVGAWAFI GLTASIVTKT LSFSAAFSAF TSEVIWLIVI SFFFARGFVK TGLGDRIATY F VKWLGKST LGLSYGLTLS EALIAPAMPS TTARAGGIFL PIIKSLSLSA GSKPNDSSSR KLGSYLIQSQ FQCAGNSSAL FL TAAAQNL LCLKLAEELG VVISNPWVSW FKAASLPAII SLLCTPLILY KLYPPETKDT PEAPGIAATK LKQMGPVTKN EWI MVGTML LAVTLWICGE TLGIPSVVAA MIGLSILLVL GVLNWDDCLS EKSAWDTLAW FAVLVGMAGQ LTNLGVVTWM SDCV AKVLQ SLSLSWPAAF GLLQAAYFFI HYLFASQTGH VGALFSAFLA MHIAAGVPGI LAALALAYNT NLFGALTHYS SGQAA VYYG AGYVDLPDVF KIGFVMATIN AIIWGVVGTF WWKFLGLY UniProtKB: Dicarboxylate transporter 2.1, chloroplastic |
-Macromolecule #2: (2S)-2-hydroxybutanedioic acid
| Macromolecule | Name: (2S)-2-hydroxybutanedioic acid / type: ligand / ID: 2 / Number of copies: 2 / Formula: LMR |
|---|---|
| Molecular weight | Theoretical: 134.087 Da |
| Chemical component information | 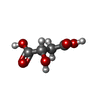 ChemComp-LMR: |
-Macromolecule #3: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 3 / Number of copies: 2 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 2 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 30 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 20 mM HEPES pH 7.4, 150 mM NaCl |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: TFS FALCON 4i (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





































