+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of THIK1 complexed with halothane | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / Ion channel / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of excitatory synapse pruning / Tandem pore domain halothane-inhibited K+ channel (THIK) / regulation of NLRP3 inflammasome complex assembly / Phase 4 - resting membrane potential / potassium ion leak channel activity / regulation of resting membrane potential / outward rectifier potassium channel activity / monoatomic ion channel complex / potassium channel activity / potassium ion transmembrane transport ...regulation of excitatory synapse pruning / Tandem pore domain halothane-inhibited K+ channel (THIK) / regulation of NLRP3 inflammasome complex assembly / Phase 4 - resting membrane potential / potassium ion leak channel activity / regulation of resting membrane potential / outward rectifier potassium channel activity / monoatomic ion channel complex / potassium channel activity / potassium ion transmembrane transport / protein heterodimerization activity / metal ion binding / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.92 Å | |||||||||
 Authors Authors | Dai Z / Yin YX | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: The structure of THIK1 complexed with halothane Authors: Dai Z / Yin YX | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_63737.map.gz emd_63737.map.gz | 54.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-63737-v30.xml emd-63737-v30.xml emd-63737.xml emd-63737.xml | 18.6 KB 18.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_63737_fsc.xml emd_63737_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_63737.png emd_63737.png | 39.3 KB | ||
| Filedesc metadata |  emd-63737.cif.gz emd-63737.cif.gz | 5.8 KB | ||
| Others |  emd_63737_additional_1.map.gz emd_63737_additional_1.map.gz emd_63737_half_map_1.map.gz emd_63737_half_map_1.map.gz emd_63737_half_map_2.map.gz emd_63737_half_map_2.map.gz | 59.7 MB 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-63737 http://ftp.pdbj.org/pub/emdb/structures/EMD-63737 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63737 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63737 | HTTPS FTP |
-Related structure data
| Related structure data |  9m9qMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_63737.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_63737.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.808 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_63737_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_63737_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_63737_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : THIK1 complexed with halothane
| Entire | Name: THIK1 complexed with halothane |
|---|---|
| Components |
|
-Supramolecule #1: THIK1 complexed with halothane
| Supramolecule | Name: THIK1 complexed with halothane / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Potassium channel subfamily K member 13
| Macromolecule | Name: Potassium channel subfamily K member 13 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.161113 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAGRGFSWGP GHLNEDNARF LLLAALIVLY LLGGAAVFSA LELAHERQAK QRWEERLANF SRGHNLSRDE LRGFLRHYEE ATRAGIRVD NVRPRWDFTG AFYFVGTVVS TIGFGMTTPA TVGGKIFLIF YGLVGCSSTI LFFNLFLERL ITIIAYIMKS C HQRQLRRR ...String: MAGRGFSWGP GHLNEDNARF LLLAALIVLY LLGGAAVFSA LELAHERQAK QRWEERLANF SRGHNLSRDE LRGFLRHYEE ATRAGIRVD NVRPRWDFTG AFYFVGTVVS TIGFGMTTPA TVGGKIFLIF YGLVGCSSTI LFFNLFLERL ITIIAYIMKS C HQRQLRRR GALPQESLKD AGQCEVDSLA GWKPSVYYVM LILCTASILI SCCASAMYTP IEGWSYFDSL YFCFVAFSTI GF GDLVSSQ NAHYESQGLY RFANFVFILM GVCCIYSLFN VISILIKQSL NWILRKMDSG CCPQCQRGLL RSRRNVVMPG SVR NRCNIS IETDGVAESD TDGRRLSGEM ISMKDLLAAN KASLAILQKQ LSEMANGCPH QTSTLARDNE FSGGVGAFAI MNNR LAETS GDRLEGGVEG GVEENLYFQ UniProtKB: Potassium channel subfamily K member 13 |
-Macromolecule #2: 2-BROMO-2-CHLORO-1,1,1-TRIFLUOROETHANE
| Macromolecule | Name: 2-BROMO-2-CHLORO-1,1,1-TRIFLUOROETHANE / type: ligand / ID: 2 / Number of copies: 2 / Formula: HLT |
|---|---|
| Molecular weight | Theoretical: 197.382 Da |
| Chemical component information | 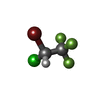 ChemComp-HLT: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.2 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 1 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































