+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Native GluA1/GluA4-CNIH3 complex in active state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AMPA receptor / native / GluA1 / GluA4 / LBD / TMD / CNIH3 / active state / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationCOPII-mediated vesicle transport / Cargo concentration in the ER / Activation of AMPA receptors / Trafficking of AMPA receptors / Trafficking of GluR2-containing AMPA receptors / Unblocking of NMDA receptors, glutamate binding and activation / axonal spine / cellular response to ammonium ion / dendritic spine membrane / long-term synaptic depression ...COPII-mediated vesicle transport / Cargo concentration in the ER / Activation of AMPA receptors / Trafficking of AMPA receptors / Trafficking of GluR2-containing AMPA receptors / Unblocking of NMDA receptors, glutamate binding and activation / axonal spine / cellular response to ammonium ion / dendritic spine membrane / long-term synaptic depression / perisynaptic space / AMPA glutamate receptor activity / negative regulation of smooth muscle cell apoptotic process / AMPA glutamate receptor complex / long-term memory / vesicle-mediated transport / synapse assembly / excitatory synapse / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / synaptic transmission, glutamatergic / receptor internalization / recycling endosome / postsynaptic density membrane / modulation of chemical synaptic transmission / synaptic vesicle membrane / dendritic spine / signaling receptor binding / neuronal cell body / synapse / dendrite / glutamatergic synapse / cell surface / endoplasmic reticulum / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.65 Å | |||||||||
 Authors Authors | Li X / Li R / Wei Y / Zhao Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2026 Journal: Cell Res / Year: 2026Title: Assembly and gating mechanism of native AMPA receptors from the cerebellum. Authors: Xiaojing Li / Renjie Li / Yiqing Wei / Jiexin Chen / Jiaojiao Zhao / Jun Zhao / Wei Wang / Na Li / Lili Wang / Tuo Hu / Yanli Dong / Yongping Zhu / Chao Wei / Long Li / Wei Zhang / Zhuo Huang / Yan Zhao /  Abstract: AMPA receptors (AMPARs) mediate the majority of fast excitatory synaptic transmission throughout the central nervous system. Calcium-permeable AMPARs and GluA4-containing receptors are critical for ...AMPA receptors (AMPARs) mediate the majority of fast excitatory synaptic transmission throughout the central nervous system. Calcium-permeable AMPARs and GluA4-containing receptors are critical for cerebellar functions, such as motor learning, associative memory, auditory processing, and synaptic plasticity. In contrast to the well-characterized, predominantly GluA2-containing AMPARs of the hippocampus and cortex, cerebellar AMPARs contain a higher proportion of GluA4 and remain poorly understood. Here, we generated a highly GluA4-specific antibody. Using this antibody in combination with antibodies specifically recognizing GluA1 and GluA2, we purified native AMPARs and determined the subunit compositions of both calcium-impermeable and calcium-permeable native AMPARs in the cerebellum. The isolated cerebellar AMPARs that contained both GluA1 and GluA4 were calcium-permeable, with GluA4 occupying mainly the B/D positions, GluA1 occupying the A/C positions, and the complex associated primarily with cornichon 3 (CNIH3). We determined the structures of the complex in distinct functional states, including the resting, active, and desensitized states, and characterized the conformational transitions that underlie its activity. During desensitization, the receptor adopts a pseudo-4-fold configuration of the ligand-binding domain layer, which may be important for its functional properties. This study provides a blueprint for the subunit compositions of AMPARs in the cerebellum and clarifies the gating mechanism of the calcium-permeable native AMPAR-CNIH3 complex, providing significant insight into AMPAR-mediated synaptic transmission in the cerebellum. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_61252.map.gz emd_61252.map.gz | 168 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-61252-v30.xml emd-61252-v30.xml emd-61252.xml emd-61252.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_61252_fsc.xml emd_61252_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_61252.png emd_61252.png | 191.9 KB | ||
| Filedesc metadata |  emd-61252.cif.gz emd-61252.cif.gz | 7 KB | ||
| Others |  emd_61252_half_map_1.map.gz emd_61252_half_map_1.map.gz emd_61252_half_map_2.map.gz emd_61252_half_map_2.map.gz | 165.1 MB 165.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-61252 http://ftp.pdbj.org/pub/emdb/structures/EMD-61252 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-61252 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-61252 | HTTPS FTP |
-Related structure data
| Related structure data |  9j94MC  9j92C  9j93C  9j95C  9x57C  9x58C  9xjlC  9xjmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_61252.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_61252.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_61252_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_61252_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : native GluA1/GluA4-CNIH3 complex in active state
| Entire | Name: native GluA1/GluA4-CNIH3 complex in active state |
|---|---|
| Components |
|
-Supramolecule #1: native GluA1/GluA4-CNIH3 complex in active state
| Supramolecule | Name: native GluA1/GluA4-CNIH3 complex in active state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Cornichon family AMPA receptor auxiliary protein 3
| Macromolecule | Name: Cornichon family AMPA receptor auxiliary protein 3 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 18.863311 KDa |
| Sequence | String: MAFTFAAFCY MLSLVLCAAL IFFAIWHIIA FDELRTDFKS PIDQCNPVHA RERLRNIERI CFLLRKLVLP EYSIHSLFCI MFLCAQEWL TLGLNVPLLF YHFWRYFHCP ADSSELAYDP PVVMNADTLS YCQKAWCKLA FYLLSFFYYL YCMIYTLVSS UniProtKB: Cornichon family AMPA receptor auxiliary protein 3 |
-Macromolecule #2: Glutamate receptor
| Macromolecule | Name: Glutamate receptor / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 101.565953 KDa |
| Sequence | String: MQHIFAFFCT GFLGAVVGAN FPNNIQIGGL FPNQQSQEHA AFRFALSQLT EPPKLLPQID IVNISDSFEM TYRFCSQFSK GVYAIFGFY ERRTVNMLTS FCGALHVCFI TPSFPVDTSN QFVLQLRPEL QDALISIIDH YKWQKFVYIY DADRGLSVLQ K VLDTAAEK ...String: MQHIFAFFCT GFLGAVVGAN FPNNIQIGGL FPNQQSQEHA AFRFALSQLT EPPKLLPQID IVNISDSFEM TYRFCSQFSK GVYAIFGFY ERRTVNMLTS FCGALHVCFI TPSFPVDTSN QFVLQLRPEL QDALISIIDH YKWQKFVYIY DADRGLSVLQ K VLDTAAEK NWQVTAVNIL TTTEEGYRML FQDLEKKKER LVVVDCESER LNAILGQIIK LEKNGIGYHY ILANLGFMDI DL NKFKESG ANVTGFQLVN YTDTIPAKIM QQWKTSDARD HTRVDWKRPK YTSALTYDGV KVMAEAFQSL RRQRIDISRR GNA GDCLAN PAVPWGQGID IQRALQQVRF EGLTGNVQFN EKGRRTNYTL HVIEMKHDGI RKIGYWNEDD KFVPAATDAQ AGGD NSSVQ NRTYIVTTIL EDPYVMLKKN ANQFEGNDRY EGYCVELAAE IAKHVGYSYR LEIVSDGKYG ARDPDTKAWN GMVGE LVYG RADVAVAPLT ITLVREEVID FSKPFMSLGI SIMIKKPQKS KPGVFSFLDP LAYEIWMCIV FAYIGVSVVL FLVSRF SPY EWHSEEFEEG RDQTTSDQSN EFGIFNSLWF SLGAFMQQGC DISPRSLSGR IVGGVWWFFT LIIISSYTAN LAAFLTV ER MVSPIESAED LAKQTEIAYG TLEAGSTKEF FRRSKIAVFE KMWTYMKSAE PSVFVRTTEE GMIRVRKSKG KYAYLLES T MNEYIEQRKP CDTMKVGGNL DSKGYGIATP KGSALRGPVN LAVLKLSEQG VLDKLKSKWW YDKGECGSKD SGSKDKTSA LSLSNVAGVF YILIGGLGLA MLVALIEFCY KSRSESKRMK GFCLIPQQSI NEAIRTSTLP RNSGAGASGG GSGENGRVVS HDFPKSMQS IPCMSHSTGM PLGATGL UniProtKB: Glutamate receptor |
-Macromolecule #3: Glutamate receptor
| Macromolecule | Name: Glutamate receptor / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 99.389758 KDa |
| Sequence | String: MRIICRQIVL LFSGFWGLAM GAFPSSVQIG GLFIRNTDQE YTAFRLAIFL HNTSPNASEA PFNLVPHVDN IETANSFAVT NAFCSQYSR GVFAIFGLYD KRSVHTLTSF CSALHISLIT PSFPTEGESQ FVLQLRPSLR GALLSLLDHY EWNCFVFLYD T DRGYSILQ ...String: MRIICRQIVL LFSGFWGLAM GAFPSSVQIG GLFIRNTDQE YTAFRLAIFL HNTSPNASEA PFNLVPHVDN IETANSFAVT NAFCSQYSR GVFAIFGLYD KRSVHTLTSF CSALHISLIT PSFPTEGESQ FVLQLRPSLR GALLSLLDHY EWNCFVFLYD T DRGYSILQ AIMEKAGQNG WHVSAICVEN FNDVSYRQLL EELDRRQEKK FVIDCEIERL QNILEQIVSV GKHVKGYHYI IA NLGFKDI SLERFIHGGA NVTGFQLVDF NTPMVTKLMD RWKKLDQREY PGSETPPKYT SALTYDGVLV MAETFRNLRR QKI DISRRG NAGDCLANPA APWGQGIDME RTLKQVQIQG LTGNVQFDHY GRRVNYTMDV FELKSTGPRK VGYWNDMDKL VLIQ DVPTL GNDTAAIENR TVVVTTIMES PYVMYKKNHE MFEGNDKYEG YCVDLASEIA KHIGIKYKIA IVPDGKYGAR DADTK IWNG MVGELVYGKA EIAIAPLTIT LVREEVIDFS KPFMSLGISI MIKKPQKSKP GVFSFLDPLA YEIWMCIVFA YIGVSV VLF LVSRFSPYEW HTEEPEDGKE GPSDQPPNEF GIFNSLWFSL GAFMQQGCDI SPRSLSGRIV GGVWWFFTLI IISSYTA NL AAFLTVERMV SPIESAEDLA KQTEIAYGTL DSGSTKEFFR RSKIAVYEKM WTYMRSAEPS VFTRTTAEGV ARVRKSKG K FAFLLESTMN EYIEQRKPCD TMKVGGNLDS KGYGVATPKG SSLRTPVNLA VLKLSEAGVL DKLKNKWWYD KGECGPKDS GSKDKTSALS LSNVAGVFYI LVGGLGLAML VALIEFCYKS RAEAKRMKVA KSAQTFNPTS SQNTQNLATY REGYNVYGTE SIKI UniProtKB: Glutamate receptor |
-Macromolecule #4: GAMMA-L-GLUTAMIC ACID
| Macromolecule | Name: GAMMA-L-GLUTAMIC ACID / type: ligand / ID: 4 / Number of copies: 4 / Formula: GGL |
|---|---|
| Molecular weight | Theoretical: 147.129 Da |
| Chemical component information | 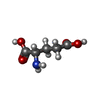 ChemComp-GGL: |
-Macromolecule #5: (3~{R})-3-[(1~{S},2~{S},4~{S})-2-bicyclo[2.2.1]hept-5-enyl]-6-chl...
| Macromolecule | Name: (3~{R})-3-[(1~{S},2~{S},4~{S})-2-bicyclo[2.2.1]hept-5-enyl]-6-chloranyl-1,1-bis(oxidanylidene)-3,4-dihydro-2~{H}-1$l^{6},2,4-benzothiadiazine-7-sulfonamide type: ligand / ID: 5 / Number of copies: 4 / Formula: A1EA4 |
|---|---|
| Molecular weight | Theoretical: 389.878 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOCONTINUUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





































