+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Receptor | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Receptor / Cryo-EM / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein localization to cytoskeleton / positive regulation of heat generation / 1-(4-iodo-2,5-dimethoxyphenyl)propan-2-amine binding / Gq/11-coupled serotonin receptor activity / positive regulation of phosphatidylinositol biosynthetic process / G protein-coupled serotonin receptor signaling pathway / G protein-coupled serotonin receptor complex / neurofilament / cell body fiber / phospholipase C-activating serotonin receptor signaling pathway ...protein localization to cytoskeleton / positive regulation of heat generation / 1-(4-iodo-2,5-dimethoxyphenyl)propan-2-amine binding / Gq/11-coupled serotonin receptor activity / positive regulation of phosphatidylinositol biosynthetic process / G protein-coupled serotonin receptor signaling pathway / G protein-coupled serotonin receptor complex / neurofilament / cell body fiber / phospholipase C-activating serotonin receptor signaling pathway / artery smooth muscle contraction / positive regulation of cytokine production involved in immune response / serotonin receptor activity / Serotonin receptors / G protein-coupled serotonin receptor activity / serotonin receptor signaling pathway / sensitization / urinary bladder smooth muscle contraction / neurotransmitter receptor activity / serotonin binding / positive regulation of platelet aggregation / negative regulation of synaptic transmission, glutamatergic / positive regulation of DNA biosynthetic process / temperature homeostasis / detection of temperature stimulus involved in sensory perception of pain / regulation of dopamine secretion / negative regulation of potassium ion transport / protein tyrosine kinase activator activity / positive regulation of vasoconstriction / positive regulation of execution phase of apoptosis / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / detection of mechanical stimulus involved in sensory perception of pain / positive regulation of fat cell differentiation / behavioral response to cocaine / release of sequestered calcium ion into cytosol / presynaptic modulation of chemical synaptic transmission / positive regulation of glycolytic process / dendritic shaft / glycolytic process / caveola / memory / Olfactory Signaling Pathway / Activation of the phototransduction cascade / intracellular calcium ion homeostasis / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / positive regulation of inflammatory response / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / ADP signalling through P2Y purinoceptor 12 / photoreceptor disc membrane / Glucagon-type ligand receptors / Sensory perception of sweet, bitter, and umami (glutamate) taste / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / cellular response to catecholamine stimulus / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / positive regulation of neuron apoptotic process / G-protein beta-subunit binding / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / extracellular vesicle / sensory perception of taste / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / positive regulation of cytosolic calcium ion concentration / retina development in camera-type eye / virus receptor activity / GTPase binding / presynaptic membrane / cytoplasmic vesicle / Ca2+ pathway / fibroblast proliferation / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / chemical synaptic transmission / Ras protein signal transduction / postsynaptic membrane / Extra-nuclear estrogen signaling / positive regulation of ERK1 and ERK2 cascade Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.84 Å | |||||||||
 Authors Authors | Min H / Kim K | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of Receptor Authors: Min H / Kim K | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_61218.map.gz emd_61218.map.gz | 450.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-61218-v30.xml emd-61218-v30.xml emd-61218.xml emd-61218.xml | 19.9 KB 19.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_61218.png emd_61218.png | 29.5 KB | ||
| Filedesc metadata |  emd-61218.cif.gz emd-61218.cif.gz | 6.5 KB | ||
| Others |  emd_61218_half_map_1.map.gz emd_61218_half_map_1.map.gz emd_61218_half_map_2.map.gz emd_61218_half_map_2.map.gz | 442.9 MB 442.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-61218 http://ftp.pdbj.org/pub/emdb/structures/EMD-61218 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-61218 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-61218 | HTTPS FTP |
-Related structure data
| Related structure data |  9j87MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_61218.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_61218.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.664 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_61218_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_61218_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of serotonin 2A with G alpha q, beta and gamma
| Entire | Name: Ternary complex of serotonin 2A with G alpha q, beta and gamma |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of serotonin 2A with G alpha q, beta and gamma
| Supramolecule | Name: Ternary complex of serotonin 2A with G alpha q, beta and gamma type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: 5-hydroxytryptamine receptor 2A
| Macromolecule | Name: 5-hydroxytryptamine receptor 2A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.619594 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSGSGLSLLH LQEKNWSALL TAVVIILTIA GNILVIMAVS LEKKLQNATN YFLMSLAIAD MLLGFLVMPV SMLTILYGYR WPLPSKLCA VWIYLDVLFS TASIMHLCAI SLDRYVAIQN PIHHSRFNSR TKAFLKIIAV WTISVGISMP IPVFGLQDDS K VFKEGSCL ...String: GSGSGLSLLH LQEKNWSALL TAVVIILTIA GNILVIMAVS LEKKLQNATN YFLMSLAIAD MLLGFLVMPV SMLTILYGYR WPLPSKLCA VWIYLDVLFS TASIMHLCAI SLDRYVAIQN PIHHSRFNSR TKAFLKIIAV WTISVGISMP IPVFGLQDDS K VFKEGSCL LADDNFVLIG SFVSFFIPLT IMVITYFLTI KSLQKEATLC VSDLGTRAKL ASFSFLPQSS LSSEKLFQRS IH REPGSYT GRRTMQSISN EQKACKVLGI VFFLFVVMWC PFFITNIMAV ICKESCNEDV IGALLNVFVW IGYLSSAVNP LVY TLFNKT YRSAFSRYIQ CQYKENKK UniProtKB: 5-hydroxytryptamine receptor 2A |
-Macromolecule #2: G protein subunit q (Gi2-mini-Gq chimera)
| Macromolecule | Name: G protein subunit q (Gi2-mini-Gq chimera) / type: protein_or_peptide / ID: 2 Details: For chain B (Gi2-mini-Gq chimeric), since this engineered protein is not listed in Uniprot, we referred to the following publication for its preparation.(PDB 6WHA) Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.084832 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSTVSAEDK AAAERSKMID KNLREDGEKA RRTLRLLLLG ADNSGKSTIV KQMRILHGGS GGSGGTSGIF ETKFQVDKVN FHMFDVGGQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA ...String: MGSTVSAEDK AAAERSKMID KNLREDGEKA RRTLRLLLLG ADNSGKSTIV KQMRILHGGS GGSGGTSGIF ETKFQVDKVN FHMFDVGGQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA RYTTPEDATP EPGEDPRVTR AKYFIRKEFV DISTASGDGR HICYPHFTCA VDTENARRIF NDCKDIILQM NL REYNLV |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.285734 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW ...String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW DIETGQQTTT FTGHTGDVMS LSLAPDTRLF VSGACDASAK LWDVREGMCR QTFTGHESDI NAICFFPNGN AF ATGSDDA TCRLFDLRAD QELMTYSHDN IICGITSVSF SKSGRLLLAG YDDFNCNVWD ALKADRAGVL AGHDNRVSCL GVT DDGMAV ATGSWDSFLK IWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: single-chain variable fragment 16 (scFv16)
| Macromolecule | Name: single-chain variable fragment 16 (scFv16) / type: protein_or_peptide / ID: 5 Details: The cis peptide bond between TYR E 223 and PRO E 224 is intentional and reflects the structural constraints of the protein. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.679721 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAA |
-Macromolecule #6: 2-(5-methoxy-1H-indol-3-yl)ethanamine
| Macromolecule | Name: 2-(5-methoxy-1H-indol-3-yl)ethanamine / type: ligand / ID: 6 / Number of copies: 1 / Formula: F5U |
|---|---|
| Molecular weight | Theoretical: 190.242 Da |
| Chemical component information | 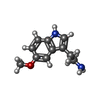 ChemComp-F5U: |
-Macromolecule #7: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 7 / Number of copies: 2 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 1.42 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































