+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Helical Structure of dITP-activated KomBC complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacterial immunity / NADase / Filament / CryoEM. / IMMUNE SYSTEM | |||||||||
| Biological species |  | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Hao F / Eddie T / Kai S / Bin W / Min L | |||||||||
| Funding support |  Singapore, 1 items Singapore, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM Helical Structure of dITP-activated KomBC complex Authors: Hao F / Eddie T / Bin W / Min L | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60415.map.gz emd_60415.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60415-v30.xml emd-60415-v30.xml emd-60415.xml emd-60415.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_60415_fsc.xml emd_60415_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  emd_60415.png emd_60415.png | 137.7 KB | ||
| Filedesc metadata |  emd-60415.cif.gz emd-60415.cif.gz | 5.8 KB | ||
| Others |  emd_60415_half_map_1.map.gz emd_60415_half_map_1.map.gz emd_60415_half_map_2.map.gz emd_60415_half_map_2.map.gz | 48.4 MB 48.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60415 http://ftp.pdbj.org/pub/emdb/structures/EMD-60415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60415 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60415 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_60415.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60415.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_60415_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_60415_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Filament of octomeric HAM-SIR2 with dITP
| Entire | Name: Filament of octomeric HAM-SIR2 with dITP |
|---|---|
| Components |
|
-Supramolecule #1: Filament of octomeric HAM-SIR2 with dITP
| Supramolecule | Name: Filament of octomeric HAM-SIR2 with dITP / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 824 kDa/nm |
-Macromolecule #1: KomB, HAM-like protein, Non-canonical purine NTP pyrophosphatase
| Macromolecule | Name: KomB, HAM-like protein, Non-canonical purine NTP pyrophosphatase type: protein_or_peptide / ID: 1 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 20.929131 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNIRFITRNR HKIKEINKIL SGTGVVVLAS EHSIDEIQTE NVHALIKDKL LKAFKLVGRP VFVEHTGLYI ESLNGFPGGL TQIFWDKLQ ADKFSQLLGT SENPRLVAKT IIGYCDSMKI YIFEGETQGT ISPVPKGPRD FQWDCIFIPD GESETFAEMG D RKNEISMR KKAFDKFKEY LLEGGK |
-Macromolecule #2: KomC, SIR2 domain protein, NADase
| Macromolecule | Name: KomC, SIR2 domain protein, NADase / type: protein_or_peptide / ID: 2 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 30.655889 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEQLLADYKK GNVILFVGAG VSMNLGLPSW SQLVDHIATE LGYDPDIYRT FGSALELAEY YKLKKGKIGP LRSWMDRMWH SSDIDINKS KVHEYIAKAN FPIIYTTNYD RWIETALSNY GKEYIKISSV SDIAKIDNNK TQIIKFHGDF DDDSSIVLDE T SYFQRLEF ...String: MEQLLADYKK GNVILFVGAG VSMNLGLPSW SQLVDHIATE LGYDPDIYRT FGSALELAEY YKLKKGKIGP LRSWMDRMWH SSDIDINKS KVHEYIAKAN FPIIYTTNYD RWIETALSNY GKEYIKISSV SDIAKIDNNK TQIIKFHGDF DDDSSIVLDE T SYFQRLEF ETPLDIKFRS DVLGKSVLFI GYSLSDINIR LLFYKLSKLW KEQKLEEAQP KSYIFLPRPN PIQEEILEQW RI GMISSEN DNPGESLEEF LKNFVLV |
-Macromolecule #3: 2'-deoxyinosine 5'-triphosphate
| Macromolecule | Name: 2'-deoxyinosine 5'-triphosphate / type: ligand / ID: 3 / Number of copies: 16 / Formula: Y43 |
|---|---|
| Molecular weight | Theoretical: 492.166 Da |
| Chemical component information | 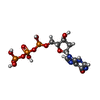 ChemComp-Y43: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 16 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





































