[English] 日本語
 Yorodumi
Yorodumi- EMDB-60399: Cryo-EM structure of GPR119-Gs Complex with small molecule agonis... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of GPR119-Gs Complex with small molecule agonist GSK-1292263 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPR119 / GSK-1292263 / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationSynthesis, secretion, and inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) / phosphatidylcholine binding / insulin secretion / regulation of metabolic process / PKA activation in glucagon signalling / hair follicle placode formation / developmental growth / D1 dopamine receptor binding / intracellular transport / vascular endothelial cell response to laminar fluid shear stress ...Synthesis, secretion, and inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) / phosphatidylcholine binding / insulin secretion / regulation of metabolic process / PKA activation in glucagon signalling / hair follicle placode formation / developmental growth / D1 dopamine receptor binding / intracellular transport / vascular endothelial cell response to laminar fluid shear stress / renal water homeostasis / Hedgehog 'off' state / adenylate cyclase-activating adrenergic receptor signaling pathway / activation of adenylate cyclase activity / regulation of insulin secretion / cellular response to glucagon stimulus / adenylate cyclase activator activity / trans-Golgi network membrane / negative regulation of inflammatory response to antigenic stimulus / G protein-coupled receptor activity / bone development / G-protein beta/gamma-subunit complex binding / platelet aggregation / cognition / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / adenylate cyclase-activating G protein-coupled receptor signaling pathway / G-protein activation / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through CDC42 / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / cellular response to catecholamine stimulus / ADORA2B mediated anti-inflammatory cytokines production / ADP signalling through P2Y purinoceptor 1 / G beta:gamma signalling through PI3Kgamma / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / adenylate cyclase-activating dopamine receptor signaling pathway / sensory perception of smell / GPER1 signaling / Inactivation, recovery and regulation of the phototransduction cascade / cellular response to prostaglandin E stimulus / G-protein beta-subunit binding / heterotrimeric G-protein complex / G alpha (12/13) signalling events / sensory perception of taste / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / positive regulation of cold-induced thermogenesis / G protein activity / GTPase binding / retina development in camera-type eye / Ca2+ pathway / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / fibroblast proliferation / G alpha (i) signalling events / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell population proliferation / receptor complex / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase activity / synapse / protein-containing complex binding / GTP binding / signal transduction / extracellular exosome / metal ion binding / membrane / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
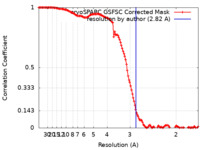
| Method | single particle reconstruction / cryo EM / Resolution: 2.82 Å | |||||||||
 Authors Authors | Wong TS / Xiong TT / Zeng ZC / Gan SY / Qiu C / Du Y | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To be published Journal: To be publishedTitle: Cryo-EM structure of GPR119-Gs complex Authors: Wong TS / Xiong TT / Zeng ZC | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60399.map.gz emd_60399.map.gz | 203.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60399-v30.xml emd-60399-v30.xml emd-60399.xml emd-60399.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_60399_fsc.xml emd_60399_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_60399.png emd_60399.png | 55.2 KB | ||
| Filedesc metadata |  emd-60399.cif.gz emd-60399.cif.gz | 6.3 KB | ||
| Others |  emd_60399_half_map_1.map.gz emd_60399_half_map_1.map.gz emd_60399_half_map_2.map.gz emd_60399_half_map_2.map.gz | 200.7 MB 200.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60399 http://ftp.pdbj.org/pub/emdb/structures/EMD-60399 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60399 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60399 | HTTPS FTP |
-Related structure data
| Related structure data |  8zrkMC  8zr5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_60399.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60399.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_60399_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_60399_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of GPR119-Gs Complex with small molecule agonis...
| Entire | Name: Cryo-EM structure of GPR119-Gs Complex with small molecule agonist GSK-1292263 |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of GPR119-Gs Complex with small molecule agonis...
| Supramolecule | Name: Cryo-EM structure of GPR119-Gs Complex with small molecule agonist GSK-1292263 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.699434 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKNTIVKQM RILHVNGFNG EGGEEDPQAA RSNSDGEKA TKVQDIKNNL KEAIETIVAA MSNLVPPVEL ANPENQFRVD YILSVMNVPD FDFPPEFYEH AKALWEDEGV R ACYERSNE ...String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKNTIVKQM RILHVNGFNG EGGEEDPQAA RSNSDGEKA TKVQDIKNNL KEAIETIVAA MSNLVPPVEL ANPENQFRVD YILSVMNVPD FDFPPEFYEH AKALWEDEGV R ACYERSNE YQLIDCAQYF LDKIDVIKQA DYVPSDQDLL RCRVLTSGIF ETKFQVDKVN FHMFDVGAQR DERRKWIQCF ND VTAIIFV VASSSYNMVI REDNQTNRLQ AALKLFDSIW NNKWLRDTSV ILFLNKQDLL AEKVLAGKSK IEDYFPEFAR YTT PEDATP EPGEDPRVTR AKYFIRDEFL RISTASGDGR HYCYPHFTCS VDTENIRRVF NDCRDIIQRM HLRQYELL UniProtKB: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.581844 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHHHH HLEVLFQGPG SSGSELDQLR QEAEQLKNQI RDARKACADA TLSQITNNID PVGRIQMRTR RTLRGHLAKI YAMHWGTDS RLLVSASQDG KLIIWDSYTT NKVHAIPLRS SWVMTCAYAP SGNYVACGGL DNICSIYNLK TREGNVRVSR E LAGHTGYL ...String: MHHHHHHHHH HLEVLFQGPG SSGSELDQLR QEAEQLKNQI RDARKACADA TLSQITNNID PVGRIQMRTR RTLRGHLAKI YAMHWGTDS RLLVSASQDG KLIIWDSYTT NKVHAIPLRS SWVMTCAYAP SGNYVACGGL DNICSIYNLK TREGNVRVSR E LAGHTGYL SCCRFLDDNQ IVTSSGDTTC ALWDIETGQQ TTTFTGHTGD VMSLSLAPDT RLFVSGACDA SAKLWDVREG MC RQTFTGH ESDINAICFF PNGNAFATGS DDATCRLFDL RADQELMTYS HDNIICGITS VSFSKSGRLL LAGYDDFNCN VWD ALKADR AGVLAGHDNR VSCLGVTDDG MAVATGSWDS FLKIWNVSGW RLFKKISVSG WRLFKKIS UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: nanobody35
| Macromolecule | Name: nanobody35 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 17.057271 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKYLLPTAAA GLLLLAAQPA MAMQVQLQES GGGLVQPGGS LRLSCAASGF TFSNYKMNWV RQAPGKGLEW VSDISQSGAS ISYTGSVKG RFTISRDNAK NTLYLQMNSL KPEDTAVYYC ARCPAPFTRD CFDVTSTTYA YRGQGTQVTV SSHHHHHH |
-Macromolecule #5: Glucose-dependent insulinotropic receptor
| Macromolecule | Name: Glucose-dependent insulinotropic receptor / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.762062 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDKLEVLFQ GPGSMESSFS FGVILAVLAS LIIATNTLVA VAVLLLIHKN DGVSLCFTLN LAVADTLIG VAISGLLTDQ LSSPSRPTQK TLCSLRMAFV TSSAAASVLT VMLITFDRYL AIKQPFRYLK IMSGFVAGAC I AGLWLVSY ...String: MKTIIALSYI FCLVFADYKD DDDKLEVLFQ GPGSMESSFS FGVILAVLAS LIIATNTLVA VAVLLLIHKN DGVSLCFTLN LAVADTLIG VAISGLLTDQ LSSPSRPTQK TLCSLRMAFV TSSAAASVLT VMLITFDRYL AIKQPFRYLK IMSGFVAGAC I AGLWLVSY LIGFLPLGIP MFQQTAYKGQ CSFFAVFHPH FVLTLSCVGF FPAMLLFVFF YCDMLKIASM HSQQIRKMEH AG AMAGGYR SPRTPSDFKA LRTVSVLIGS FALSWTPFLI TGIVQVACQE CHLYLVLERY LWLLGVGNSL LNPLIYAYWQ KEV RLQLYH MALGVKKVLT SFLLFLSARN CGPERPRESS CHIVTISSSE FDG UniProtKB: Glucose-dependent insulinotropic receptor |
-Macromolecule #6: GSK-1292263
| Macromolecule | Name: GSK-1292263 / type: ligand / ID: 6 / Number of copies: 1 / Formula: A1D8Y |
|---|---|
| Molecular weight | Theoretical: 456.558 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Sugar embedding | Material: vitreous ice |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 51.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: OTHER |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.4000000000000001 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)