+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human 26S immunoproteasome | |||||||||
 Map data Map data | Locally filtered, globally sharpened map using a local resolution estimation computed from the half-maps. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | proteasome / immunoproteasome / HYDROLASE | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Saratov GA / Baymukhametov TN / Konevega AL / Kudriaeva AA / Belogurov AA | |||||||||
| Funding support |  Russian Federation, 1 items Russian Federation, 1 items
| |||||||||
 Citation Citation |  Journal: Russ.J.Bioorganic Chem. / Year: 2024 Journal: Russ.J.Bioorganic Chem. / Year: 2024Title: The 3.6 Angstroms cryo-EM structure of the outer heptameric alpha-ring of human 26S immunoproteasome in the preactivation state. Authors: Saratov GA / Baymukhametov TN / Konevega AL / Kudriaeva AA / Belogurov AA | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60139.map.gz emd_60139.map.gz | 3.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60139-v30.xml emd-60139-v30.xml emd-60139.xml emd-60139.xml | 53.3 KB 53.3 KB | Display Display |  EMDB header EMDB header |
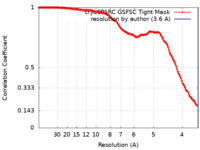
| FSC (resolution estimation) |  emd_60139_fsc.xml emd_60139_fsc.xml | 14.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_60139.png emd_60139.png | 85.5 KB | ||
| Masks |  emd_60139_msk_1.map emd_60139_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-60139.cif.gz emd-60139.cif.gz | 13.7 KB | ||
| Others |  emd_60139_additional_1.map.gz emd_60139_additional_1.map.gz emd_60139_additional_2.map.gz emd_60139_additional_2.map.gz emd_60139_additional_3.map.gz emd_60139_additional_3.map.gz emd_60139_half_map_1.map.gz emd_60139_half_map_1.map.gz emd_60139_half_map_2.map.gz emd_60139_half_map_2.map.gz | 3.4 MB 3.4 MB 226.5 MB 7.5 MB 7.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60139 http://ftp.pdbj.org/pub/emdb/structures/EMD-60139 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60139 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60139 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_60139.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60139.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally filtered, globally sharpened map using a local resolution estimation computed from the half-maps. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.8 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_60139_msk_1.map emd_60139_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: The map post-processed by deepEMhancer using a default...
| File | emd_60139_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The map post-processed by deepEMhancer using a default deep learning model 'tightTarget' (with mask). | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: The map post-processed by deepEMhancer using a default...
| File | emd_60139_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The map post-processed by deepEMhancer using a default deep learning model 'tightTarget' (without mask). | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unfiltered and unsharpened map obtained using cryoSPARC's 3D...
| File | emd_60139_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered and unsharpened map obtained using cryoSPARC's 3D flexible refinement. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: The half-map masked by a relatively wide (15...
| File | emd_60139_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The half-map masked by a relatively wide (15 angstrom dilation in a 256 px box) solvent soft mask due to the peculiarities of the cryoSPARC's 3D Flex Reconstruct procedure. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: The half-map masked by a relatively wide (15...
| File | emd_60139_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The half-map masked by a relatively wide (15 angstrom dilation in a 256 px box) solvent soft mask due to the peculiarities of the cryoSPARC's 3D Flex Reconstruct procedure. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Human 26S immunoproteasome
+Supramolecule #1: Human 26S immunoproteasome
+Macromolecule #1: Proteasome subunit alpha type-1
+Macromolecule #2: Proteasome subunit alpha type-2
+Macromolecule #3: Proteasome subunit alpha type-3
+Macromolecule #4: Proteasome subunit alpha type-4
+Macromolecule #5: Proteasome subunit alpha type-5
+Macromolecule #6: Proteasome subunit alpha type-6
+Macromolecule #7: Proteasome subunit alpha type-7
+Macromolecule #8: Proteasome subunit beta type-1
+Macromolecule #9: Proteasome subunit beta type-2
+Macromolecule #10: Proteasome subunit beta type-3
+Macromolecule #11: Proteasome subunit beta type-4
+Macromolecule #12: Proteasome subunit beta type-8
+Macromolecule #13: Proteasome subunit beta type-9
+Macromolecule #14: Proteasome subunit beta type-10
+Macromolecule #15: 26S proteasome regulatory subunit 4
+Macromolecule #16: 26S proteasome regulatory subunit 6A
+Macromolecule #17: 26S proteasome regulatory subunit 6B
+Macromolecule #18: 26S proteasome regulatory subunit 7
+Macromolecule #19: 26S proteasome regulatory subunit 8
+Macromolecule #20: 26S proteasome regulatory subunit 10B
+Macromolecule #21: 26S proteasome non-ATPase regulatory subunit 1
+Macromolecule #22: 26S proteasome non-ATPase regulatory subunit 2
+Macromolecule #23: 26S proteasome non-ATPase regulatory subunit 3
+Macromolecule #24: 26S proteasome non-ATPase regulatory subunit 4
+Macromolecule #25: 26S proteasome non-ATPase regulatory subunit 6
+Macromolecule #26: 26S proteasome non-ATPase regulatory subunit 7
+Macromolecule #27: 26S proteasome non-ATPase regulatory subunit 8
+Macromolecule #28: 26S proteasome non-ATPase regulatory subunit 9
+Macromolecule #29: 26S proteasome non-ATPase regulatory subunit 11
+Macromolecule #30: 26S proteasome non-ATPase regulatory subunit 12
+Macromolecule #31: 26S proteasome non-ATPase regulatory subunit 13
+Macromolecule #32: 26S proteasome non-ATPase regulatory subunit 14 (His)x6 TEV prote...
+Macromolecule #33: Proteasomal ubiquitin receptor ADRM1
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.9 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 30mM Tris-HCl, 5mM MgCl2, 1mM ATP, 1mM TCEP |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GRAPHENE OXIDE / Mesh: 400 / Support film - Material: GRAPHENE OXIDE / Support film - topology: HOLEY ARRAY / Details: The grid was not pretreated. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: Microscope was modified with a Cs corrector (CEOS GmbH, Germany). |
| Details | Preliminary grid screening was performed manually. |
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 1-20 / Number grids imaged: 1 / Number real images: 1130 / Average exposure time: 3.0 sec. / Average electron dose: 60.0 e/Å2 Details: Images were collected in movie-mode at 20 frames per second. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 77778 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 37000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)