[English] 日本語
 Yorodumi
Yorodumi- EMDB-5699: CryoEM structure of EGFR-specific virus (derived from sindbis virus) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5699 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of EGFR-specific virus (derived from sindbis virus) | |||||||||
 Map data Map data | CryoEM structure of EGFR-specific virus derived from sindbis virus | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | EGFR-specific virus / sindbis virus | |||||||||
| Biological species | synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 12.2 Å | |||||||||
 Authors Authors | Dai H / Liu Z / Jiang W / Kuhn RJ | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2013 Journal: J Virol / Year: 2013Title: Directed evolution of a virus exclusively utilizing human epidermal growth factor receptor as the entry receptor. Authors: Hong-Sheng Dai / Zheng Liu / Wen Jiang / Richard J Kuhn Abstract: Rational design and directed evolution are powerful tools to generate and improve protein function; however, their uses are mostly limited to enzyme and antibody engineering. Here we describe a ...Rational design and directed evolution are powerful tools to generate and improve protein function; however, their uses are mostly limited to enzyme and antibody engineering. Here we describe a directed-evolution strategy, named the tandem selection and enrichment system (TSES), and its use in generating virus with exclusive specificity for a particular cellular receptor. In TSES, evolving viruses are sequentially and iteratively transferred between two different host cells, one for selection of receptor specificity and the other for enrichment of the fittest virus. By combining rational design and TSES, we generated human epidermal growth factor receptor (EGFR)-specific virus 1 (ESV1). ESV1 has the backbone of Sindbis virus (SINV) and displays an EGF domain engrafted onto structural protein E2 after residue Pro192, together with eight amino acid changes stabilizing the E2-EGF chimera. ESV1 uses EGFR to initiate infection and has lost the capacity to interact with all known SINV receptors. A 12.2-Å cryoelectron microscopic (cryoEM) reconstruction of ESV1 reveals that the E2-EGF fusion adopts a fixed conformation, with EGF sitting at the top of the E2 spike; The EGFR binding interface faces outward, and the EGF domain completely masks SINV receptor binding. The cryoEM structure of ESV1 explains the desirable properties of ESV1 and provides insights for its further modification. TSES expands the scope of directed evolution and can be easily extended to other targeting molecules and viral systems. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5699.map.gz emd_5699.map.gz | 33.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5699-v30.xml emd-5699-v30.xml emd-5699.xml emd-5699.xml | 11.2 KB 11.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5699.png emd_5699.png | 5.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5699 http://ftp.pdbj.org/pub/emdb/structures/EMD-5699 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5699 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5699 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5699.map.gz / Format: CCP4 / Size: 126.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5699.map.gz / Format: CCP4 / Size: 126.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of EGFR-specific virus derived from sindbis virus | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.24 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
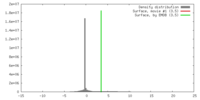
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : EGFR-specific virus (derived from sindbis virus)
| Entire | Name: EGFR-specific virus (derived from sindbis virus) |
|---|---|
| Components |
|
-Supramolecule #1000: EGFR-specific virus (derived from sindbis virus)
| Supramolecule | Name: EGFR-specific virus (derived from sindbis virus) / type: sample / ID: 1000 / Details: The sample was monodisperse. / Oligomeric state: icosahedral / Number unique components: 1 |
|---|
-Supramolecule #1: synthetic construct
| Supramolecule | Name: synthetic construct / type: virus / ID: 1 / Name.synonym: EGFR-specific Sindbis virus Details: EGF domain was engrafted onto an icosahedral virus scaffold, and directed evolution was conducted to stabilize the chimeric virus. NCBI-ID: 32630 / Sci species name: synthetic construct / Sci species strain: EGFR-specific no.1 / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No / Syn species name: EGFR-specific Sindbis virus |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Virus shell | Shell ID: 1 / Name: E1 and E2-EGF / Diameter: 375.00 Å / T number (triangulation number): 4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 50 mM Tris, 200 mM NaCl, 1 mM EDTA |
| Grid | Details: 400 mesh copper grid with thin carbon support, glow discharged in air |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 70 % / Chamber temperature: 97.15 K / Instrument: GATAN CRYOPLUNGE 3 / Method: Blot for 6 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Temperature | Min: 93.15 K / Max: 100.15 K / Average: 97 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 100,000 times magnification |
| Date | Oct 12, 2011 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 3.24 µm / Number real images: 166 / Average electron dose: 20 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 39190 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 38000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
- Image processing
Image processing
| CTF correction | Details: each particle |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 12.2 Å / Resolution method: OTHER / Software - Name: jspr, EMAN2, EMAN / Details: Final maps were calculated from 2290 particles. / Number images used: 2290 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: veda |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
-Atomic model buiding 2
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)