+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5639 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron tomography reconstruction of native HIV-1 core | |||||||||
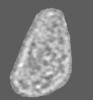
 Map data Map data | The figure shows the simulated density of all-atom HIV-1 capsid model (3J3Y) overlaid with a slice of HIV-1 core tomogram. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cryo-electron tomography / HIV-1 core | |||||||||
| Function / homology |  Function and homology information Function and homology informationviral budding via host ESCRT complex / host multivesicular body / viral nucleocapsid / viral translational frameshifting / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity / RNA binding / zinc ion binding ...viral budding via host ESCRT complex / host multivesicular body / viral nucleocapsid / viral translational frameshifting / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity / RNA binding / zinc ion binding / identical protein binding / membrane Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Zhao G / Perilla JR / Yufenyuy E / Meng X / Chen B / Ning J / Ahn J / Gronenborn AM / Schulten K / Aiken C / Zhang P | |||||||||
 Citation Citation |  Journal: Nature / Year: 2013 Journal: Nature / Year: 2013Title: Mature HIV-1 capsid structure by cryo-electron microscopy and all-atom molecular dynamics. Authors: Gongpu Zhao / Juan R Perilla / Ernest L Yufenyuy / Xin Meng / Bo Chen / Jiying Ning / Jinwoo Ahn / Angela M Gronenborn / Klaus Schulten / Christopher Aiken / Peijun Zhang /  Abstract: Retroviral capsid proteins are conserved structurally but assemble into different morphologies. The mature human immunodeficiency virus-1 (HIV-1) capsid is best described by a 'fullerene cone' model, ...Retroviral capsid proteins are conserved structurally but assemble into different morphologies. The mature human immunodeficiency virus-1 (HIV-1) capsid is best described by a 'fullerene cone' model, in which hexamers of the capsid protein are linked to form a hexagonal surface lattice that is closed by incorporating 12 capsid-protein pentamers. HIV-1 capsid protein contains an amino-terminal domain (NTD) comprising seven α-helices and a β-hairpin, a carboxy-terminal domain (CTD) comprising four α-helices, and a flexible linker with a 310-helix connecting the two structural domains. Structures of the capsid-protein assembly units have been determined by X-ray crystallography; however, structural information regarding the assembled capsid and the contacts between the assembly units is incomplete. Here we report the cryo-electron microscopy structure of a tubular HIV-1 capsid-protein assembly at 8 Å resolution and the three-dimensional structure of a native HIV-1 core by cryo-electron tomography. The structure of the tubular assembly shows, at the three-fold interface, a three-helix bundle with critical hydrophobic interactions. Mutagenesis studies confirm that hydrophobic residues in the centre of the three-helix bundle are crucial for capsid assembly and stability, and for viral infectivity. The cryo-electron-microscopy structures enable modelling by large-scale molecular dynamics simulation, resulting in all-atom models for the hexamer-of-hexamer and pentamer-of-hexamer elements as well as for the entire capsid. Incorporation of pentamers results in closer trimer contacts and induces acute surface curvature. The complete atomic HIV-1 capsid model provides a platform for further studies of capsid function and for targeted pharmacological intervention. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5639.map.gz emd_5639.map.gz | 3.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5639-v30.xml emd-5639-v30.xml emd-5639.xml emd-5639.xml | 11.3 KB 11.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5639_1.jpg emd_5639_1.jpg emd_5639_2.jpg emd_5639_2.jpg | 62.6 KB 108.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5639 http://ftp.pdbj.org/pub/emdb/structures/EMD-5639 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5639 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5639 | HTTPS FTP |
-Validation report
| Summary document |  emd_5639_validation.pdf.gz emd_5639_validation.pdf.gz | 254.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5639_full_validation.pdf.gz emd_5639_full_validation.pdf.gz | 254.2 KB | Display | |
| Data in XML |  emd_5639_validation.xml.gz emd_5639_validation.xml.gz | 4.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5639 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5639 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5639 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5639 | HTTPS FTP |
-Related structure data
| Related structure data |  3j3qMC  3j3yMC  5582C  3j34C  3j4fC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5639.map.gz / Format: CCP4 / Size: 17.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5639.map.gz / Format: CCP4 / Size: 17.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The figure shows the simulated density of all-atom HIV-1 capsid model (3J3Y) overlaid with a slice of HIV-1 core tomogram. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 6.25 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : HIV-1 core with A14C/E45C cross-linked capsid protein
| Entire | Name: HIV-1 core with A14C/E45C cross-linked capsid protein |
|---|---|
| Components |
|
-Supramolecule #1000: HIV-1 core with A14C/E45C cross-linked capsid protein
| Supramolecule | Name: HIV-1 core with A14C/E45C cross-linked capsid protein / type: sample / ID: 1000 / Details: The sample was purified from HIV-1 virions / Number unique components: 1 |
|---|
-Supramolecule #1: Human immunodeficiency virus 1
| Supramolecule | Name: Human immunodeficiency virus 1 / type: virus / ID: 1 / Name.synonym: HIV-1 Details: HIV-1 core was isolated from virions carrying A14C/E45C mutation in capsid protein. NCBI-ID: 11676 / Sci species name: Human immunodeficiency virus 1 / Sci species strain: R9 / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No / Syn species name: HIV-1 |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.011 mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 10 mM Tris-HCl, 100 mM NaCl, 1 mM EDTA |
| Grid | Details: Quantifoil R2/2 200 mesh holey carbon copper grids |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 70 % / Chamber temperature: 90 K / Instrument: HOMEMADE PLUNGER Method: Purified HIV-1 A14C/E45C cores (3 uL) were applied to the carbon side of glow-discharged, perforated R2/2 Quantifoil grids and quickly mixed with 3 uL of a 15 nM fiducial gold bead solution ...Method: Purified HIV-1 A14C/E45C cores (3 uL) were applied to the carbon side of glow-discharged, perforated R2/2 Quantifoil grids and quickly mixed with 3 uL of a 15 nM fiducial gold bead solution before plunge-freezing. |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Temperature | Min: 80 K / Max: 85 K / Average: 82 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 115,000 times magnification |
| Date | Sep 11, 2010 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 53 / Average electron dose: 120 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 8.0 µm / Nominal defocus min: 8.0 µm / Nominal magnification: 39000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC / Tilt series - Axis1 - Min angle: -70 ° / Tilt series - Axis1 - Max angle: 66 ° / Tilt series - Axis1 - Angle increment: 2 ° |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: OTHER / Software - Name: tomo3d Details: The raw tomogram was denoised using the nonlinear anisotropic diffusion edge-enhancing algorithm available in IMOD. The core density was segmented out using Volume Tracer in UCSF Chimera. Number images used: 53 |
|---|
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: MD |
| Details | The model was built using hexamer of hexamers (PDB entry 3J34) and pentamer of hexamers (computer-based MD model available upon request). |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-3j3q:  PDB-3j3y: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)