[English] 日本語
 Yorodumi
Yorodumi- EMDB-5502: Dissecting the in vivo assembly of the 30S ribosomal subunit reve... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5502 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dissecting the in vivo assembly of the 30S ribosomal subunit reveals the role of RimM | |||||||||
 Map data Map data | The map has been normalized to N(0,1) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribosome biogenesis / 30S subunit assembly / RimM | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 13.1 Å | |||||||||
 Authors Authors | Guo Q / Goto S / Chen Y / Muto A / Himeno H / Deng H / Lei J / Gao N | |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2013 Journal: Nucleic Acids Res / Year: 2013Title: Dissecting the in vivo assembly of the 30S ribosomal subunit reveals the role of RimM and general features of the assembly process. Authors: Qiang Guo / Simon Goto / Yuling Chen / Boya Feng / Yanji Xu / Akira Muto / Hyouta Himeno / Haiteng Deng / Jianlin Lei / Ning Gao /  Abstract: Ribosome biogenesis is a tightly regulated, multi-stepped process. The assembly of ribosomal subunits is a central step of the complex biogenesis process, involving nearly 30 protein factors in vivo ...Ribosome biogenesis is a tightly regulated, multi-stepped process. The assembly of ribosomal subunits is a central step of the complex biogenesis process, involving nearly 30 protein factors in vivo in bacteria. Although the assembly process has been extensively studied in vitro for over 40 years, very limited information is known for the in vivo process and specific roles of assembly factors. Such an example is ribosome maturation factor M (RimM), a factor involved in the late-stage assembly of the 30S subunit. Here, we combined quantitative mass spectrometry and cryo-electron microscopy to characterize the in vivo 30S assembly intermediates isolated from mutant Escherichia coli strains with genes for assembly factors deleted. Our compositional and structural data show that the assembly of the 3'-domain of the 30S subunit is severely delayed in these intermediates, featured with highly underrepresented 3'-domain proteins and large conformational difference compared with the mature 30S subunit. Further analysis indicates that RimM functions not only to promote the assembly of a few 3'-domain proteins but also to stabilize the rRNA tertiary structure. More importantly, this study reveals intriguing similarities and dissimilarities between the in vitro and the in vivo assembly pathways, suggesting that they are in general similar but with subtle differences. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5502.map.gz emd_5502.map.gz | 6.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5502-v30.xml emd-5502-v30.xml emd-5502.xml emd-5502.xml | 10 KB 10 KB | Display Display |  EMDB header EMDB header |
| Images |  400_5502.gif 400_5502.gif 80_5502.gif 80_5502.gif | 41.2 KB 3.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5502 http://ftp.pdbj.org/pub/emdb/structures/EMD-5502 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5502 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5502 | HTTPS FTP |
-Related structure data
| Related structure data |  3j2aMC  5500C  5501C  5503C  5504C  5506C  5507C  5508C  5509C  5510C  3j28C  3j29C  3j2bC  3j2cC  3j2dC  3j2eC  3j2fC  3j2gC  3j2hC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5502.map.gz / Format: CCP4 / Size: 7.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5502.map.gz / Format: CCP4 / Size: 7.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
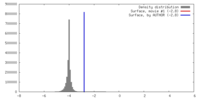
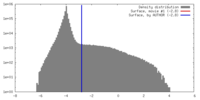
| Annotation | The map has been normalized to N(0,1) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.76 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Immature ribosomal small subunit from rimm gene deleted E.coli strain
| Entire | Name: Immature ribosomal small subunit from rimm gene deleted E.coli strain |
|---|---|
| Components |
|
-Supramolecule #1000: Immature ribosomal small subunit from rimm gene deleted E.coli strain
| Supramolecule | Name: Immature ribosomal small subunit from rimm gene deleted E.coli strain type: sample / ID: 1000 / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 800 KDa / Theoretical: 800 KDa |
-Supramolecule #1: Small subunit from rimm gene deleted E.coli strain
| Supramolecule | Name: Small subunit from rimm gene deleted E.coli strain / type: complex / ID: 1 / Name.synonym: immature 30S / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-prokaryote: SSU 30S |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 800 KDa / Theoretical: 800 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 150mM NH4Cl,10mM Tris-HCL,10mM MgCl2 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV / Method: Blot for 1 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Jan 1, 2012 |
| Image recording | Category: CCD / Film or detector model: GENERIC GATAN (4k x 4k) / Average electron dose: 20 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.3 µm / Nominal magnification: 80000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | This is a classification volume (No. 3) using ML3D methods. |
|---|---|
| CTF correction | Details: Weiner filter |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 13.1 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Spider / Number images used: 30262 |
-Atomic model buiding 1
| Initial model | PDB ID:  3ofa Chain - Chain ID: A |
|---|---|
| Software | Name: MDFF |
| Details | Protocol: Initial local fitting was done using Chimera and then MDFF was used for flexible fitting. ref: Trabuco, L.G., Villa, E., Mitra, K., Frank, J. and Schulten, K. (2008) Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure, 16, 673-683 |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: Cross-correlation |
| Output model |  PDB-3j2a: |
 Movie
Movie Controller
Controller






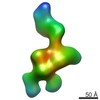


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)