+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5370 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of full-length NSF in the ADP-AlFx state | |||||||||
 Map data Map data | NSF hexamer in the ADP-AlFx state | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | membrane fusion / AAA+ ATPase | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 9.2 Å | |||||||||
 Authors Authors | Chang LF / Chen S / Liu CC / Pan X / Jiang J / Bai XC / Xie X / Wang HW / Sui SF | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2012 Journal: Nat Struct Mol Biol / Year: 2012Title: Structural characterization of full-length NSF and 20S particles. Authors: Lei-Fu Chang / Song Chen / Cui-Cui Liu / Xijiang Pan / Jiansen Jiang / Xiao-Chen Bai / Xin Xie / Hong-Wei Wang / Sen-Fang Sui /  Abstract: The 20S particle, which is composed of the N-ethylmaleimide-sensitive factor (NSF), soluble NSF attachment proteins (SNAPs) and the SNAP receptor (SNARE) complex, has an essential role in ...The 20S particle, which is composed of the N-ethylmaleimide-sensitive factor (NSF), soluble NSF attachment proteins (SNAPs) and the SNAP receptor (SNARE) complex, has an essential role in intracellular vesicle fusion events. Using single-particle cryo-EM and negative stain EM, we reconstructed four related three-dimensional structures: Chinese hamster NSF hexamer in the ATPγS, ADP-AlFx and ADP states, and the 20S particle. These structures reveal a parallel arrangement between the D1 and D2 domains of the hexameric NSF and characterize the nucleotide-dependent conformational changes in NSF. The structure of the 20S particle shows that it holds the SNARE complex at two interaction interfaces around the C terminus and N-terminal half of the SNARE complex, respectively. These findings provide insight into the molecular mechanism underlying disassembly of the SNARE complex by NSF. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5370.map.gz emd_5370.map.gz | 6.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5370-v30.xml emd-5370-v30.xml emd-5370.xml emd-5370.xml | 7.3 KB 7.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5370_1.jpg emd_5370_1.jpg | 181.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5370 http://ftp.pdbj.org/pub/emdb/structures/EMD-5370 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5370 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5370 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5370.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5370.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | NSF hexamer in the ADP-AlFx state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.8 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
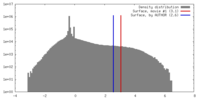
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : N-ethylmaleimide-sensitive factor (NSF)
| Entire | Name: N-ethylmaleimide-sensitive factor (NSF) |
|---|---|
| Components |
|
-Supramolecule #1000: N-ethylmaleimide-sensitive factor (NSF)
| Supramolecule | Name: N-ethylmaleimide-sensitive factor (NSF) / type: sample / ID: 1000 / Details: The sample was freshly purified. / Oligomeric state: homohexamer / Number unique components: 1 |
|---|
-Macromolecule #1: N-ethylmaleimide-sensitive factor
| Macromolecule | Name: N-ethylmaleimide-sensitive factor / type: protein_or_peptide / ID: 1 / Name.synonym: NSF / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Sep 29, 2010 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder: CT3500 / Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 9.2 Å / Resolution method: FSC 0.5 CUT-OFF |
|---|
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)