[English] 日本語
 Yorodumi
Yorodumi- EMDB-5344: Human Mediator-RNA polymerase II-TFIIF assembly in the absence of... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5344 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Mediator-RNA polymerase II-TFIIF assembly in the absence of activator | |||||||||
 Map data Map data | activator-free Mediator bound to pol II & TFIIF | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transcription / Mediator / RNA polymerase II / TFIIF / activator | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 32.0 Å | |||||||||
 Authors Authors | Bernecky C / Taatjes DJ | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2012 Journal: J Mol Biol / Year: 2012Title: Activator-mediator binding stabilizes RNA polymerase II orientation within the human mediator-RNA polymerase II-TFIIF assembly. Authors: Carrie Bernecky / Dylan J Taatjes /  Abstract: The human Mediator complex controls RNA polymerase II (pol II) function in ways that remain incompletely understood. Activator-Mediator binding alters Mediator structure, and these activator-induced ...The human Mediator complex controls RNA polymerase II (pol II) function in ways that remain incompletely understood. Activator-Mediator binding alters Mediator structure, and these activator-induced structural shifts appear to play key roles in regulating transcription. A recent cryo-electron microscopy (EM) analysis revealed that pol II adopted a stable orientation within a Mediator-pol II-TFIIF assembly in which Mediator was bound to the activation domain of viral protein 16 (VP16). Whereas TFIIF was shown to be important for orienting pol II within this assembly, the potential role of the activator was not assessed. To determine how activator binding might affect pol II orientation, we isolated human Mediator-pol II-TFIIF complexes in which Mediator was not bound to an activator. Cryo-EM analysis of this assembly, coupled with pol II crystal structure docking, revealed that pol II binds Mediator at the same general location; however, in contrast to VP16-bound Mediator, pol II does not appear to stably orient in the absence of an activator. Variability in pol II orientation might be important mechanistically, perhaps to enable sense and antisense transcription at human promoters. Because Mediator interacts extensively with pol II, these results suggest that Mediator structural shifts induced by activator binding help stably orient pol II prior to transcription initiation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5344.map.gz emd_5344.map.gz | 14.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5344-v30.xml emd-5344-v30.xml emd-5344.xml emd-5344.xml | 11.8 KB 11.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5344_1.png emd_5344_1.png | 45.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5344 http://ftp.pdbj.org/pub/emdb/structures/EMD-5344 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5344 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5344 | HTTPS FTP |
-Validation report
| Summary document |  emd_5344_validation.pdf.gz emd_5344_validation.pdf.gz | 78.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5344_full_validation.pdf.gz emd_5344_full_validation.pdf.gz | 77.3 KB | Display | |
| Data in XML |  emd_5344_validation.xml.gz emd_5344_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5344 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5344 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5344 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5344 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5344.map.gz / Format: CCP4 / Size: 15.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5344.map.gz / Format: CCP4 / Size: 15.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | activator-free Mediator bound to pol II & TFIIF | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.22 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
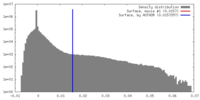
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Assembly of human Mediator, RNA polymerase II, and TFIIF
| Entire | Name: Assembly of human Mediator, RNA polymerase II, and TFIIF |
|---|---|
| Components |
|
-Supramolecule #1000: Assembly of human Mediator, RNA polymerase II, and TFIIF
| Supramolecule | Name: Assembly of human Mediator, RNA polymerase II, and TFIIF type: sample / ID: 1000 Oligomeric state: one Mediator complex binds one pol II-TFIIF Number unique components: 3 |
|---|---|
| Molecular weight | Theoretical: 1.9 MDa |
-Macromolecule #1: core Mediator
| Macromolecule | Name: core Mediator / type: protein_or_peptide / ID: 1 / Name.synonym: Mediator Details: affinity purified using an antibody to the MED26 subunit Oligomeric state: 26 subunit complex / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: human / Cell: HeLa / Organelle: Nucleus Homo sapiens (human) / synonym: human / Cell: HeLa / Organelle: Nucleus |
| Molecular weight | Theoretical: 1.2 MDa |
-Macromolecule #2: RNA polymerase II
| Macromolecule | Name: RNA polymerase II / type: protein_or_peptide / ID: 2 / Name.synonym: pol II / Details: unphosphorylated / Oligomeric state: 12-subunit complex / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Cell: HeLa / Organelle: Nucleus Homo sapiens (human) / synonym: Human / Cell: HeLa / Organelle: Nucleus |
| Molecular weight | Theoretical: 520 KDa |
-Macromolecule #3: TFIIF
| Macromolecule | Name: TFIIF / type: protein_or_peptide / ID: 3 / Name.synonym: TFIIF / Details: RAP74 and RAP30 expressed separately then combined / Oligomeric state: Dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 10 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 Details: 20 mM HEPES, 0.10 mM EDTA, 150 mM KCl, 0.02% NP-40, 35% glycerol |
|---|---|
| Staining | Type: NEGATIVE Details: grids with adsorbed protein washed 3x with buffer containing 5% trehalose, 20 mM HEPES, 100 mM KCl, and 0.10 mM EDTA, then subject to cryo-negative staining in a saturated solution (1.2M) of ...Details: grids with adsorbed protein washed 3x with buffer containing 5% trehalose, 20 mM HEPES, 100 mM KCl, and 0.10 mM EDTA, then subject to cryo-negative staining in a saturated solution (1.2M) of ammonium molybdate (pH 7.5) |
| Grid | Details: thin carbon-coated holey carbon 400 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 90 K / Instrument: OTHER Method: blot for 2 seconds, dry for 3 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 12.7 µm / Number real images: 139 / Average electron dose: 15 e/Å2 / Od range: 1 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 29000 |
| Sample stage | Specimen holder: side entry / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The particles were selected interactively at the computer terminal. |
|---|---|
| CTF correction | Details: each micrograph |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 32.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Spider / Number images used: 5489 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: Situs |
| Details | Protocol: rigid body. the TFIIS chain S was removed before fitting |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: contour-based Laplacian correlation |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)