[English] 日本語
 Yorodumi
Yorodumi- EMDB-5171: Structure of Fusarium poae virus 1 shows conserved and variable e... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5171 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Fusarium poae virus 1 shows conserved and variable elements of partitivirus capsids and evolutionary links to picobirnaviruses | |||||||||
 Map data Map data | map of fpv1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Capsid protein Cryoelectron microscopy dsRNA virus Electron cryomicroscopy Fungal virus Virus structure | |||||||||
| Biological species | fpv1 (unknown) | |||||||||
| Method | single particle reconstruction / cryo EM | |||||||||
 Authors Authors | Tang J / Ochoa WF / Li H / Havens WM / Nibert ML / Ghabrial SA / Baker TS | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2010 Journal: J Struct Biol / Year: 2010Title: Structure of Fusarium poae virus 1 shows conserved and variable elements of partitivirus capsids and evolutionary relationships to picobirnavirus. Authors: Jinghua Tang / Wendy F Ochoa / Hua Li / Wendy M Havens / Max L Nibert / Said A Ghabrial / Timothy S Baker /  Abstract: Filamentous fungus Fusarium poae is a worldwide cause of the economically important disease Fusarium head blight of cereal grains. The fungus is itself commonly infected with a bisegmented dsRNA ...Filamentous fungus Fusarium poae is a worldwide cause of the economically important disease Fusarium head blight of cereal grains. The fungus is itself commonly infected with a bisegmented dsRNA virus from the family Partitiviridae. For this study, we determined the structure of partitivirus Fusarium poae virus 1 (FpV1) to a resolution of 5.6Å or better by electron cryomicroscopy and three-dimensional image reconstruction. The main structural features of FpV1 are consistent with those of two other fungal partitiviruses for which high-resolution structures have been recently reported. These shared features include a 120-subunit T=1 capsid comprising 60 quasisymmetrical capsid protein dimers with both shell and protruding domains. Distinguishing features are evident throughout the FpV1 capsid, however, consistent with its more massive subunits and its greater phylogenetic divergence relative to the other two structurally characterized partitiviruses. These results broaden our understanding of conserved and variable elements of fungal partitivirus structure, as well as that of vertebrate picobirnavirus, and support the suggestion that a phylogenetic subcluster of partitiviruses closely related to FpV1 should constitute a separate taxonomic genus. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5171.map.gz emd_5171.map.gz | 29.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5171-v30.xml emd-5171-v30.xml emd-5171.xml emd-5171.xml | 7.3 KB 7.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5171_1.jpg emd_5171_1.jpg | 71.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5171 http://ftp.pdbj.org/pub/emdb/structures/EMD-5171 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5171 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5171 | HTTPS FTP |
-Validation report
| Summary document |  emd_5171_validation.pdf.gz emd_5171_validation.pdf.gz | 77.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5171_full_validation.pdf.gz emd_5171_full_validation.pdf.gz | 76.2 KB | Display | |
| Data in XML |  emd_5171_validation.xml.gz emd_5171_validation.xml.gz | 498 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5171 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5171 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5171 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5171 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5171.map.gz / Format: CCP4 / Size: 74.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5171.map.gz / Format: CCP4 / Size: 74.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map of fpv1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.95 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
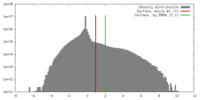
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : fpv1
| Entire | Name: fpv1 |
|---|---|
| Components |
|
-Supramolecule #1000: fpv1
| Supramolecule | Name: fpv1 / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: fpv1
| Supramolecule | Name: fpv1 / type: virus / ID: 1 / Name.synonym: fpv1 / Sci species name: fpv1 / Database: NCBI / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: fpv1 |
|---|---|
| Host (natural) | Organism:  Fungi (eukaryote) / synonym: FUNGI Fungi (eukaryote) / synonym: FUNGI |
| Virus shell | Shell ID: 1 / Diameter: 420 Å / T number (triangulation number): 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Date | Jul 21, 2006 |
| Image recording | Digitization - Sampling interval: 1.95 µm / Number real images: 240 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)