+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Rea1 D2915A-R2976A-D3042A mutant ATP conformation III | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AAA+ protein / ribosome maturation / TRANSLOCASE | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.9 Å | |||||||||
 Authors Authors | Busselez J / Schmidt H | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Remodelling of Rea1 linker domain drives the removal of assembly factors from pre-ribosomal particles. Authors: Johan Busselez / Geraldine Koenig / Carine Dominique / Torben Klos / Deepika Velayudhan / Piotr Sosnowski / Nils Marechal / Corinne Crucifix / Hugo Gizardin-Fredon / Sarah Cianferani / ...Authors: Johan Busselez / Geraldine Koenig / Carine Dominique / Torben Klos / Deepika Velayudhan / Piotr Sosnowski / Nils Marechal / Corinne Crucifix / Hugo Gizardin-Fredon / Sarah Cianferani / Benjamin Albert / Yves Henry / Anthony K Henras / Helgo Schmidt /   Abstract: The ribosome maturation factor Rea1 (or Midasin) catalyses the removal of assembly factors from large ribosomal subunit precursors and promotes their export from the nucleus to the cytosol. Rea1 ...The ribosome maturation factor Rea1 (or Midasin) catalyses the removal of assembly factors from large ribosomal subunit precursors and promotes their export from the nucleus to the cytosol. Rea1 consists of nearly 5000 amino-acid residues and belongs to the AAA+ protein family. It consists of a ring of six AAA+ domains from which the ≈1700 amino-acid residue linker emerges that is subdivided into stem, middle and top domains. A flexible and unstructured D/E rich region connects the linker top to a MIDAS (metal ion dependent adhesion site) domain, which is able to bind the assembly factor substrates. Despite its key importance for ribosome maturation, the mechanism driving assembly factor removal by Rea1 is still poorly understood. Here we demonstrate that the Rea1 linker is essential for assembly factor removal. It rotates and swings towards the AAA+ ring following a complex remodelling scheme involving nucleotide independent as well as nucleotide dependent steps. ATP-hydrolysis is required to engage the linker with the AAA+ ring and ultimately with the AAA+ ring docked MIDAS domain. The interaction between the linker top and the MIDAS domain allows direct force transmission for assembly factor removal. | |||||||||
| History |
|
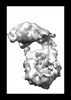
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_50818.map.gz emd_50818.map.gz | 6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-50818-v30.xml emd-50818-v30.xml emd-50818.xml emd-50818.xml | 13.3 KB 13.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_50818.png emd_50818.png | 58.4 KB | ||
| Filedesc metadata |  emd-50818.cif.gz emd-50818.cif.gz | 4 KB | ||
| Others |  emd_50818_half_map_1.map.gz emd_50818_half_map_1.map.gz emd_50818_half_map_2.map.gz emd_50818_half_map_2.map.gz | 6 MB 6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-50818 http://ftp.pdbj.org/pub/emdb/structures/EMD-50818 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50818 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50818 | HTTPS FTP |
-Validation report
| Summary document |  emd_50818_validation.pdf.gz emd_50818_validation.pdf.gz | 596.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_50818_full_validation.pdf.gz emd_50818_full_validation.pdf.gz | 596.2 KB | Display | |
| Data in XML |  emd_50818_validation.xml.gz emd_50818_validation.xml.gz | 8.4 KB | Display | |
| Data in CIF |  emd_50818_validation.cif.gz emd_50818_validation.cif.gz | 10 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50818 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50818 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50818 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50818 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_50818.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_50818.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.44 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_50818_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_50818_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Rea1 monomer
| Entire | Name: Rea1 monomer |
|---|---|
| Components |
|
-Supramolecule #1: Rea1 monomer
| Supramolecule | Name: Rea1 monomer / type: organelle_or_cellular_component / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 500 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK II |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 IS (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 6.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 240000 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)