+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | in situ 70S Myxococcus xanthus ribosome | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | in situ Subtomogram averaging / Myxococcus xanthus / RIBOSOME | |||||||||
| Biological species |  Myxococcus xanthus (bacteria) Myxococcus xanthus (bacteria) | |||||||||
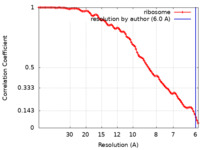
| Method | subtomogram averaging / cryo EM / Resolution: 6.0 Å | |||||||||
 Authors Authors | Xu P / Schumacher D / Liu C / Dickmanns M / Beck F / Plitzko J / Baumeister W / Sogaard-Andersen L | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2025 Journal: Proc Natl Acad Sci U S A / Year: 2025Title: In situ architecture of a nucleoid-associated biomolecular co-condensate that regulates bacterial cell division. Authors: Peng Xu / Dominik Schumacher / Chuan Liu / Andrea Harms / Marcel Dickmanns / Florian Beck / Jürgen M Plitzko / Wolfgang Baumeister / Lotte Søgaard-Andersen /  Abstract: In most bacteria, cell division depends on the tubulin-homolog FtsZ that polymerizes in a GTP-dependent manner to form the cytokinetic Z-ring at the future division site. Subsequently, the Z-ring ...In most bacteria, cell division depends on the tubulin-homolog FtsZ that polymerizes in a GTP-dependent manner to form the cytokinetic Z-ring at the future division site. Subsequently, the Z-ring recruits, directly or indirectly, all other proteins of the divisome complex that executes cytokinesis. A critical step in this process is the precise positioning of the Z-ring at the future division site. While the divisome proteins are generally conserved, the regulatory systems that position the Z-ring are more diverse. However, these systems have in common that they modulate FtsZ polymerization. In PomX, PomY, and PomZ form precisely one MDa-sized, nonstoichiometric, nucleoid-associated assembly that spatiotemporally guides Z-ring formation. Here, using cryo-correlative light and electron microscopy together with in situ cryoelectron tomography, we determine the PomXYZ assembly's architecture at close-to-live conditions. PomX forms a porous meshwork of randomly intertwined filaments. Templated by this meshwork, the phase-separating PomY protein forms a biomolecular condensate that compacts and bends the PomX filaments, resulting in the formation of a selective PomXYZ co-condensate that is associated to the nucleoid by PomZ. These studies reveal a hitherto undescribed supramolecular structure and provide a framework for understanding how a nonstoichiometric co-condensate forms, maintains number control, and nucleates GTP-dependent FtsZ polymerization to precisely regulate cell division. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_50504.map.gz emd_50504.map.gz | 73.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-50504-v30.xml emd-50504-v30.xml emd-50504.xml emd-50504.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_50504_fsc.xml emd_50504_fsc.xml | 9.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_50504.png emd_50504.png | 71.7 KB | ||
| Filedesc metadata |  emd-50504.cif.gz emd-50504.cif.gz | 4.6 KB | ||
| Others |  emd_50504_half_map_1.map.gz emd_50504_half_map_1.map.gz emd_50504_half_map_2.map.gz emd_50504_half_map_2.map.gz | 40.3 MB 40.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-50504 http://ftp.pdbj.org/pub/emdb/structures/EMD-50504 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50504 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50504 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_50504.map.gz / Format: CCP4 / Size: 78.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_50504.map.gz / Format: CCP4 / Size: 78.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.93 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_50504_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_50504_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : in situ subtomo averaging of Myxococcus xanthus ribosome
| Entire | Name: in situ subtomo averaging of Myxococcus xanthus ribosome |
|---|---|
| Components |
|
-Supramolecule #1: in situ subtomo averaging of Myxococcus xanthus ribosome
| Supramolecule | Name: in situ subtomo averaging of Myxococcus xanthus ribosome type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Myxococcus xanthus (bacteria) Myxococcus xanthus (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
| Details | M. xanthus was grown in 1% CTT broth (1% (w/v) Bacto Casitone, 10mM Tris-HCl pH 8.0, 1mM K2HPO4/KH2PO4 pH 7.6, 8mM MgSO4) to OD600~=1.8 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 5.0 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)