+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4975 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Haemonchus galactose containing glycoprotein complex | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Multi-protease complex / hydrolase | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationprotein processing / metalloendopeptidase activity / aspartic-type endopeptidase activity / lysosome / proteolysis / extracellular region / metal ion binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Haemonchus contortus (barber pole worm) Haemonchus contortus (barber pole worm) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | ||||||||||||
 Authors Authors | Scarff CA / Thompson RF | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2020 Journal: PLoS Pathog / Year: 2020Title: Structure of the protective nematode protease complex H-gal-GP and its conservation across roundworm parasites. Authors: Charlotte A Scarff / Rebecca F Thompson / George F J Newlands / Alexander H Jamson / Christopher Kennaway / Vivian J da Silva / Elida M Rabelo / Chun-Feng Song / John Trinick / W David Smith ...Authors: Charlotte A Scarff / Rebecca F Thompson / George F J Newlands / Alexander H Jamson / Christopher Kennaway / Vivian J da Silva / Elida M Rabelo / Chun-Feng Song / John Trinick / W David Smith / Stephen P Muench /   Abstract: Roundworm parasite infections are a major cause of human and livestock disease worldwide and a threat to global food security. Disease control currently relies on anthelmintic drugs to which ...Roundworm parasite infections are a major cause of human and livestock disease worldwide and a threat to global food security. Disease control currently relies on anthelmintic drugs to which roundworms are becoming increasingly resistant. An alternative approach is control by vaccination and 'hidden antigens', components of the worm gut not encountered by the infected host, have been exploited to produce Barbervax, the first commercial vaccine for a gut dwelling nematode of any host. Here we present the structure of H-gal-GP, a hidden antigen from Haemonchus contortus, the Barber's Pole worm, and a major component of Barbervax. We demonstrate its novel architecture, subunit composition and topology, flexibility and heterogeneity using cryo-electron microscopy, mass spectrometry, and modelling. Importantly, we demonstrate that complexes with the same architecture are present in other Strongylid roundworm parasites including human hookworm. This suggests a common ancestry and the potential for development of a unified hidden antigen vaccine. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4975.map.gz emd_4975.map.gz | 107.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4975-v30.xml emd-4975-v30.xml emd-4975.xml emd-4975.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4975_fsc.xml emd_4975_fsc.xml | 12.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_4975.png emd_4975.png | 120.1 KB | ||
| Masks |  emd_4975_msk_1.map emd_4975_msk_1.map | 184 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-4975.cif.gz emd-4975.cif.gz | 6.2 KB | ||
| Others |  emd_4975_additional.map.gz emd_4975_additional.map.gz emd_4975_half_map_1.map.gz emd_4975_half_map_1.map.gz emd_4975_half_map_2.map.gz emd_4975_half_map_2.map.gz | 107.3 MB 145.4 MB 145 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4975 http://ftp.pdbj.org/pub/emdb/structures/EMD-4975 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4975 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4975 | HTTPS FTP |
-Related structure data
| Related structure data |  6rowMC  4976C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4975.map.gz / Format: CCP4 / Size: 184 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4975.map.gz / Format: CCP4 / Size: 184 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.065 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_4975_msk_1.map emd_4975_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
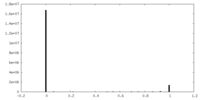
| Density Histograms |
-Additional map: #1
| File | emd_4975_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_4975_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_4975_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Haemonchus galactose containing glycoprotein complex
| Entire | Name: Haemonchus galactose containing glycoprotein complex |
|---|---|
| Components |
|
-Supramolecule #1: Haemonchus galactose containing glycoprotein complex
| Supramolecule | Name: Haemonchus galactose containing glycoprotein complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Multi-protease complex |
|---|---|
| Source (natural) | Organism:  Haemonchus contortus (barber pole worm) Haemonchus contortus (barber pole worm) |
| Molecular weight | Theoretical: 500 KDa |
-Macromolecule #1: Putative zinc metallopeptidase
| Macromolecule | Name: Putative zinc metallopeptidase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Haemonchus contortus (barber pole worm) Haemonchus contortus (barber pole worm) |
| Molecular weight | Theoretical: 86.819812 KDa |
| Sequence | String: NANRSKEWKN AANTLLFGLD ESVDPCEDFY GFTCNKFIER IDLDELGRGR FTTFSQAQLE VNSDIVKALE KVDVNDEKFS QTERITKAA FQSCVEYTLK DDRKASVNEL LQYISERFGG IPFLGQRVKG GCELYREMGR IEQQRALPTF MYTWVNVDHK N VSRNSYYI ...String: NANRSKEWKN AANTLLFGLD ESVDPCEDFY GFTCNKFIER IDLDELGRGR FTTFSQAQLE VNSDIVKALE KVDVNDEKFS QTERITKAA FQSCVEYTLK DDRKASVNEL LQYISERFGG IPFLGQRVKG GCELYREMGR IEQQRALPTF MYTWVNVDHK N VSRNSYYI SQPTLPMPRE FYVLPQFAPE LDARTKAIEN VMKAFASDIL KDPSKYDKMI KTAAREVTQI EQKIAMASWP DD ELRNHEQ QYNPYELFAP SEGKIALQDA FPNIRFGKYI AGLMETATGG VPVSPVFIGD VIVTNPAYMG FLNTLFRQTN IQM NQYPFV NYIIVHMLFE DAEHLGDKYL RIAKDADYVT YAQRKGVGIT RSGRKYSRVF DERTDARMKC VDTITTYMPY GTGY VYVNS REDRDQVVED VKQQTELIMK TFLKKMLSTL SWMQGESYRR AEKKINEMHR NYGWPKKLFG DFKNFDTIDA YHRDD YYSI LEAYNNKTDK STAFYTILNI LRRGYENRES FRRKNETADR TNFLESPASV NAWYAPELNS LTLPFGILTS PHYDLQ FPK AFNFAGSGTV GGHELVHGFD DEGVQFDYDG SLADCSVYEC GWLEQKGKNG FKDMAQCVVT QYNAQCCPAK EGNVHCA NG AHTQGENIAD LGGLQASYNA YKEYIKMKGA EEMRLPGLEK FTPNQIFWIS YGYSWCAKET QSSLVKRLLT NPHSPNSC R VNQVLQDIPS FAKDFQCALG QKMYPPAEQR CKVW UniProtKB: Putative zinc metallopeptidase |
-Macromolecule #2: Parasite pepsinogen
| Macromolecule | Name: Parasite pepsinogen / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Haemonchus contortus (barber pole worm) Haemonchus contortus (barber pole worm) |
| Molecular weight | Theoretical: 40.310469 KDa |
| Sequence | String: VFPHPIYDYQ DTEYLAKITI GAPGQSFHVV LDTGSANLWI PDNICVNGRR GACRITTCDR GLVCEVLCHD KSCCEDDVDN PDEDNPCKG KSGFDSTQST SYAKITPKKY FEIVYGTGFA KGFLGNDTVR FGEEGNNKTL VVPGTVFGQA VQIGDPFANN P INGILGLG ...String: VFPHPIYDYQ DTEYLAKITI GAPGQSFHVV LDTGSANLWI PDNICVNGRR GACRITTCDR GLVCEVLCHD KSCCEDDVDN PDEDNPCKG KSGFDSTQST SYAKITPKKY FEIVYGTGFA KGFLGNDTVR FGEEGNNKTL VVPGTVFGQA VQIGDPFANN P INGILGLG FRGLAQAGVT PPLQRAIDLK LVDPIFTVYM KQLGAKAKGQ DGGAFTYGGL DSVNCGQEIA YVDLTRPLYW QF KMEAFSA GYLSIRKGWE VISDTGTSFM GVPTAIADLV ADSYGGQYDE MFEIYTVDCN ATVTFGMTIG GKQYKIERKN LVL EEDKDS CMIAMTPLSS VGFGPQWILG APFIRQYCNI HDMRNNTIGF AEP UniProtKB: Parasite pepsinogen |
-Macromolecule #3: Cysteine Protease
| Macromolecule | Name: Cysteine Protease / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Haemonchus contortus (barber pole worm) Haemonchus contortus (barber pole worm) |
| Molecular weight | Theoretical: 28.31549 KDa |
| Sequence | String: DPDIPENYDP RLIWPNCSSL FTIPDQANCG SCWAVSTAAA ISDRLCIASK GENQVFISSA DILSCCDTCG FGCDGGITFR AWEYFATKG SVSGGHFEAP NCCRPYYFHP CGQHGNDTFY GYCPRFAQTP FCRRKCRIGF NKSYAQDRIQ GKSFYTVDYS V PAIQREIM ...String: DPDIPENYDP RLIWPNCSSL FTIPDQANCG SCWAVSTAAA ISDRLCIASK GENQVFISSA DILSCCDTCG FGCDGGITFR AWEYFATKG SVSGGHFEAP NCCRPYYFHP CGQHGNDTFY GYCPRFAQTP FCRRKCRIGF NKSYAQDRIQ GKSFYTVDYS V PAIQREIM TKGSVVGSYD VFTDFSHYKS GIYRHTGGKP DGRHAVRIIG WGKENGTDYW LIANSWHDDW GENGYFRMIR GI NDCGIEQ DITAGD |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 62.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Models were generated using Phyre2, rigid body fitted in Chimera and then flexibly fitted into the map using MDFF in VMD. |
|---|---|
| Refinement | Protocol: FLEXIBLE FIT |
| Output model |  PDB-6row: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)