+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | HsSTING with cGAMP/C53/DCA | |||||||||
 Map data Map data | HsSTING cGAMP/C53/DCA with C2 symmetry | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Innate immunity / membrane protein / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationSTING complex / STAT6-mediated induction of chemokines / serine/threonine protein kinase complex / protein localization to endoplasmic reticulum / 2',3'-cyclic GMP-AMP binding / cyclic-di-GMP binding / STING mediated induction of host immune responses / positive regulation of type I interferon-mediated signaling pathway / IRF3-mediated induction of type I IFN / proton channel activity ...STING complex / STAT6-mediated induction of chemokines / serine/threonine protein kinase complex / protein localization to endoplasmic reticulum / 2',3'-cyclic GMP-AMP binding / cyclic-di-GMP binding / STING mediated induction of host immune responses / positive regulation of type I interferon-mediated signaling pathway / IRF3-mediated induction of type I IFN / proton channel activity / reticulophagy / cGAS/STING signaling pathway / pattern recognition receptor signaling pathway / cellular response to exogenous dsRNA / cytoplasmic pattern recognition receptor signaling pathway / protein complex oligomerization / autophagosome membrane / positive regulation of macroautophagy / autophagosome assembly / cellular response to interferon-beta / positive regulation of defense response to virus by host / positive regulation of type I interferon production / endoplasmic reticulum-Golgi intermediate compartment membrane / signaling adaptor activity / activation of innate immune response / positive regulation of interferon-beta production / autophagosome / secretory granule membrane / cytoplasmic vesicle membrane / antiviral innate immune response / Regulation of innate immune responses to cytosolic DNA / protein serine/threonine kinase binding / SARS-CoV-1 activates/modulates innate immune responses / peroxisome / regulation of inflammatory response / defense response to virus / RNA polymerase II-specific DNA-binding transcription factor binding / mitochondrial outer membrane / transcription coactivator activity / endosome / cilium / ciliary basal body / Golgi membrane / innate immune response / Neutrophil degranulation / ubiquitin protein ligase binding / endoplasmic reticulum membrane / protein kinase binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses / perinuclear region of cytoplasm / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / nucleoplasm / identical protein binding / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.93 Å | |||||||||
 Authors Authors | Gharpure A / Ward AB / Lairson LL | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Microbiota-derived secondary bile acids promote STING activation and antitumor activity Authors: Yang X / Zhang X / Li W / Gharpure A / Hansel A / Tillack A / Wu C / Hsieh Y / Sulpizio A / Dikiy S / Zhao X / Forli S / Ward AB / Parker CG / Hang HC | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_49094.map.gz emd_49094.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-49094-v30.xml emd-49094-v30.xml emd-49094.xml emd-49094.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_49094.png emd_49094.png | 104.1 KB | ||
| Filedesc metadata |  emd-49094.cif.gz emd-49094.cif.gz | 6.5 KB | ||
| Others |  emd_49094_additional_1.map.gz emd_49094_additional_1.map.gz emd_49094_half_map_1.map.gz emd_49094_half_map_1.map.gz emd_49094_half_map_2.map.gz emd_49094_half_map_2.map.gz | 117.9 MB 115.7 MB 115.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-49094 http://ftp.pdbj.org/pub/emdb/structures/EMD-49094 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-49094 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-49094 | HTTPS FTP |
-Related structure data
| Related structure data |  9n7fMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_49094.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_49094.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | HsSTING cGAMP/C53/DCA with C2 symmetry | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.718 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: HsSTING cGAMP/C53/DCA with C1 symmetry
| File | emd_49094_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | HsSTING cGAMP/C53/DCA with C1 symmetry | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_49094_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_49094_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Stimulator of interferon genes
| Entire | Name: Stimulator of interferon genes |
|---|---|
| Components |
|
-Supramolecule #1: Stimulator of interferon genes
| Supramolecule | Name: Stimulator of interferon genes / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 400 KDa |
-Macromolecule #1: Stimulator of interferon genes protein
| Macromolecule | Name: Stimulator of interferon genes protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.246 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPHSSLHPSI PCPRGHGAQK AALVLLSACL VTLWGLGEPP EHTLRYLVLH LASLQLGLLL NGVCSLAEEL RHIHSRYRGS YWRTVRACL GCPLRRGALL LLSIYFYYSL PNAVGPPFTW MLALLGLSQA LNILLGLKGL APAEISAVCE KGNFNVAHGL A WSYYIGYL ...String: MPHSSLHPSI PCPRGHGAQK AALVLLSACL VTLWGLGEPP EHTLRYLVLH LASLQLGLLL NGVCSLAEEL RHIHSRYRGS YWRTVRACL GCPLRRGALL LLSIYFYYSL PNAVGPPFTW MLALLGLSQA LNILLGLKGL APAEISAVCE KGNFNVAHGL A WSYYIGYL RLILPELQAR IRTYNQHYNN LLRGAVSQRL YILLPLDCGV PDNLSMADPN IRFLDKLPQQ TGDHAGIKDR VY SNSIYEL LENGQRAGTC VLEYATPLQT LFAMSQYSQA GFSREDRLEQ AKLFCRTLED ILADAPESQN NCRLIAYQEP ADD SSFSLS QEVLRHLRQE EKEEVTVGGG SGGGSGGSAW SHPQFEK UniProtKB: Stimulator of interferon genes protein |
-Macromolecule #2: 1-[(2-chloro-6-fluorophenyl)methyl]-3,3-dimethyl-2-oxo-N-[(2,4,6-...
| Macromolecule | Name: 1-[(2-chloro-6-fluorophenyl)methyl]-3,3-dimethyl-2-oxo-N-[(2,4,6-trifluorophenyl)methyl]-2,3-dihydro-1H-indole-6-carboxamide type: ligand / ID: 2 / Number of copies: 2 / Formula: 9IM |
|---|---|
| Molecular weight | Theoretical: 490.877 Da |
| Chemical component information | 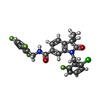 ChemComp-9IM: |
-Macromolecule #3: cGAMP
| Macromolecule | Name: cGAMP / type: ligand / ID: 3 / Number of copies: 2 / Formula: 1SY |
|---|---|
| Molecular weight | Theoretical: 674.411 Da |
| Chemical component information | 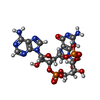 ChemComp-1SY: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 8 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: TFS FALCON 4i (4k x 4k) / Number real images: 5839 / Average exposure time: 3.2 sec. / Average electron dose: 45.12 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 20.0 µm / Illumination mode: SPOT SCAN / Imaging mode: OTHER / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 190000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































