[English] 日本語
 Yorodumi
Yorodumi- EMDB-4800: Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4800 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in extended state, 3-fold symmetrised | |||||||||
 Map data Map data | Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in extended state, 3-fold symmetrised. Scaling by factor of 2 is recommended for visualization | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Anti-feeding prophage / secretion system / AFP / contractile / VIRUS LIKE PARTICLE / baseplate | |||||||||
| Function / homology | Contractile injection system tube protein, N-terminal domain / Contractile injection system tube protein / Gp5/Type VI secretion system Vgr protein, OB-fold domain / Type VI secretion system/phage-baseplate injector OB domain / Vgr protein, OB-fold domain superfamily / Afp8 / Afp7 Function and homology information Function and homology information | |||||||||
| Biological species |  Serratia entomophila (bacteria) Serratia entomophila (bacteria) | |||||||||
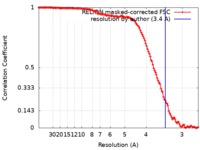
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Desfosses A | |||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2019 Journal: Nat Microbiol / Year: 2019Title: Atomic structures of an entire contractile injection system in both the extended and contracted states. Authors: Ambroise Desfosses / Hariprasad Venugopal / Tapan Joshi / Jan Felix / Matthew Jessop / Hyengseop Jeong / Jaekyung Hyun / J Bernard Heymann / Mark R H Hurst / Irina Gutsche / Alok K Mitra /       Abstract: Contractile injection systems are sophisticated multiprotein nanomachines that puncture target cell membranes. Although the number of atomic-resolution insights into contractile bacteriophage tails, ...Contractile injection systems are sophisticated multiprotein nanomachines that puncture target cell membranes. Although the number of atomic-resolution insights into contractile bacteriophage tails, bacterial type six secretion systems and R-pyocins is rapidly increasing, structural information on the contraction of bacterial phage-like protein-translocation structures directed towards eukaryotic hosts is scarce. Here, we characterize the antifeeding prophage AFP from Serratia entomophila by cryo-electron microscopy. We present the high-resolution structure of the entire AFP particle in the extended state, trace 11 protein chains de novo from the apical cap to the needle tip, describe localization variants and perform specific structural comparisons with related systems. We analyse inter-subunit interactions and highlight their universal conservation within contractile injection systems while revealing the specificities of AFP. Furthermore, we provide the structure of the AFP sheath-baseplate complex in a contracted state. This study reveals atomic details of interaction networks that accompany and define the contraction mechanism of toxin-delivery tailocins, offering a comprehensive framework for understanding their mode of action and for their possible adaptation as biocontrol agents. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4800.map.gz emd_4800.map.gz | 22.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4800-v30.xml emd-4800-v30.xml emd-4800.xml emd-4800.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4800_fsc.xml emd_4800_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_4800.png emd_4800.png | 112 KB | ||
| Filedesc metadata |  emd-4800.cif.gz emd-4800.cif.gz | 6 KB | ||
| Others |  emd_4800_half_map_1.map.gz emd_4800_half_map_1.map.gz emd_4800_half_map_2.map.gz emd_4800_half_map_2.map.gz | 140.6 MB 140.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4800 http://ftp.pdbj.org/pub/emdb/structures/EMD-4800 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4800 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4800 | HTTPS FTP |
-Related structure data
| Related structure data |  6rbkMC  4782C  4783C  4784C  4801C  4802C  4803C  4859C  4871C  4876C  6raoC  6rapC  6rbnC  6rc8C  6rglC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4800.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4800.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in extended state, 3-fold symmetrised. Scaling by factor of 2 is recommended for visualization | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: None
| File | emd_4800_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: None
| File | emd_4800_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in...
| Entire | Name: Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in extended state, 3-fold symmetrised |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in...
| Supramolecule | Name: Cryo-EM structure of the anti-feeding prophage (AFP) baseplate in extended state, 3-fold symmetrised type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: A scaling by a factor of 2 is recommended for visualisation |
|---|---|
| Source (natural) | Organism:  Serratia entomophila (bacteria) Serratia entomophila (bacteria) |
-Macromolecule #1: Afp7
| Macromolecule | Name: Afp7 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Serratia entomophila (bacteria) Serratia entomophila (bacteria) |
| Molecular weight | Theoretical: 25.259434 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSLLERGLSK LTLNAWKDRE GKIPAGSMSA MYNPETIQLD YQTRFDTEDT INTASQSNRY VISEPVGLNL TLLFDSQMPG NTTPIETQL AMLKSLCAVD AATGSPYFLR ITWGKMRWEN KGWFAGRARD LSVTYTLFDR DATPLRATVQ LSLVADESFV I QQSLKTQS ...String: MSLLERGLSK LTLNAWKDRE GKIPAGSMSA MYNPETIQLD YQTRFDTEDT INTASQSNRY VISEPVGLNL TLLFDSQMPG NTTPIETQL AMLKSLCAVD AATGSPYFLR ITWGKMRWEN KGWFAGRARD LSVTYTLFDR DATPLRATVQ LSLVADESFV I QQSLKTQS APDRALVSVP DLASLPLLAL SAGGVLASSV DYLSLAWDND LDNLDDFQTG DFLRATKGEE V UniProtKB: Afp7 |
-Macromolecule #2: Afp8
| Macromolecule | Name: Afp8 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Serratia entomophila (bacteria) Serratia entomophila (bacteria) |
| Molecular weight | Theoretical: 58.091715 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSHITLDIAG QRSTLGIRRL RVQQLINEIP LAQLELHIPT DNHGAADNAV QHEVSRFTLG VRVGIAQDNK PLFDGYLVQK KMQLKGKEW SVRLEARHAL QKLTFLPHSR VFRQQDDSTV MKGLLQSAGV KLTQESAAQL SSKHDQLLQF RLSDWQFIRS R LLSTNCWL ...String: MSHITLDIAG QRSTLGIRRL RVQQLINEIP LAQLELHIPT DNHGAADNAV QHEVSRFTLG VRVGIAQDNK PLFDGYLVQK KMQLKGKEW SVRLEARHAL QKLTFLPHSR VFRQQDDSTV MKGLLQSAGV KLTQESAAQL SSKHDQLLQF RLSDWQFIRS R LLSTNCWL LPDAASDTVV IRPLSDAAMA SRTLARDSHD YTLYEINLNF DNRFTPDSLS LQGWDIAAQR LTAAQKSPAG AF RPWKPAG KVGQSSAGRQ DYALAFSMLP EATLQTLSNS WLNYQQMTGV QGHIVLAGTR DFAPGESITL SGFGAGLDGT AML SGVNQQ FDTQYGWRSE LVIGLPASML EPAPPVRSLH IGTVAGFTAD PQHLDRIAIH LPALNLPDSL IFARLSKPWA SHAS GFCFY PEPGDEVVVG FIDSDPRYPM ILGALHNPKN TAPFPPDEKN NRKGLIVSQA DQTQALMIDT EEKTLRLMAG DNTLT LTGE GNLTMSTPNA LQLQADTLGL QADSNLSIAG KQQVEITSAK INMKK UniProtKB: Afp8 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 20.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)