+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4569 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Flagellar motor relic structure from Plesiomonas shigelloides | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Plesiomonas shigelloides ATCC 14029 (bacteria) Plesiomonas shigelloides ATCC 14029 (bacteria) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 47.0 Å | |||||||||
 Authors Authors | Ferreira JF / Beeby M | |||||||||
 Citation Citation |  Journal: PLoS Biol / Year: 2019 Journal: PLoS Biol / Year: 2019Title: γ-proteobacteria eject their polar flagella under nutrient depletion, retaining flagellar motor relic structures. Authors: Josie L Ferreira / Forson Z Gao / Florian M Rossmann / Andrea Nans / Susanne Brenzinger / Rohola Hosseini / Amanda Wilson / Ariane Briegel / Kai M Thormann / Peter B Rosenthal / Morgan Beeby /    Abstract: Bacteria switch only intermittently to motile planktonic lifestyles under favorable conditions. Under chronic nutrient deprivation, however, bacteria orchestrate a switch to stationary phase, ...Bacteria switch only intermittently to motile planktonic lifestyles under favorable conditions. Under chronic nutrient deprivation, however, bacteria orchestrate a switch to stationary phase, conserving energy by altering metabolism and stopping motility. About two-thirds of bacteria use flagella to swim, but how bacteria deactivate this large molecular machine remains unclear. Here, we describe the previously unreported ejection of polar motors by γ-proteobacteria. We show that these bacteria eject their flagella at the base of the flagellar hook when nutrients are depleted, leaving a relic of a former flagellar motor in the outer membrane. Subtomogram averages of the full motor and relic reveal that this is an active process, as a plug protein appears in the relic, likely to prevent leakage across their outer membrane; furthermore, we show that ejection is triggered only under nutritional depletion and is independent of the filament as a possible mechanosensor. We show that filament ejection is a widespread phenomenon demonstrated by the appearance of relic structures in diverse γ-proteobacteria including Plesiomonas shigelloides, Vibrio cholerae, Vibrio fischeri, Shewanella putrefaciens, and Pseudomonas aeruginosa. While the molecular details remain to be determined, our results demonstrate a novel mechanism for bacteria to halt costly motility when nutrients become scarce. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4569.map.gz emd_4569.map.gz | 53.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4569-v30.xml emd-4569-v30.xml emd-4569.xml emd-4569.xml | 8.7 KB 8.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4569.png emd_4569.png | 60.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4569 http://ftp.pdbj.org/pub/emdb/structures/EMD-4569 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4569 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4569 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4569.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4569.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
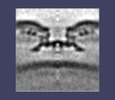
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.713 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Flagellar relic structure from Plesiomonas shigelloides
| Entire | Name: Flagellar relic structure from Plesiomonas shigelloides |
|---|---|
| Components |
|
-Supramolecule #1: Flagellar relic structure from Plesiomonas shigelloides
| Supramolecule | Name: Flagellar relic structure from Plesiomonas shigelloides type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Plesiomonas shigelloides ATCC 14029 (bacteria) / Location in cell: Outer membrane Plesiomonas shigelloides ATCC 14029 (bacteria) / Location in cell: Outer membrane |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: Cells grown in LB |
|---|---|
| Grid | Model: UltrAuFoil / Material: GOLD / Mesh: 200 |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average exposure time: 2.0 sec. / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C13 (13 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 47.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: PEET / Number subtomograms used: 124 |
|---|---|
| Extraction | Number tomograms: 89 / Number images used: 124 / Software - Name:  IMOD IMOD |
| CTF correction | Software - Name: TOMOCTF |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















