+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The Structure of Spiroplasma Virus 4 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Microviridae / bacteriophage / capsid / Spiroplasma virus 4 / SpV4 / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationT=1 icosahedral viral capsid / viral capsid / host cell cytoplasm / structural molecule activity / DNA binding Similarity search - Function | |||||||||
| Biological species |  Spiromicrovirus SpV4 Spiromicrovirus SpV4 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.52 Å | |||||||||
 Authors Authors | Mietzsch M / McKenna R | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Viruses / Year: 2024 Journal: Viruses / Year: 2024Title: The Structure of : Exploring the Capsid Diversity of the . Authors: Mario Mietzsch / Shweta Kailasan / Antonette Bennett / Paul Chipman / Bentley Fane / Juha T Huiskonen / Ian N Clarke / Robert McKenna /    Abstract: (SpV4) is a bacteriophage of the , which packages circular ssDNA within non-enveloped T = 1 icosahedral capsids. It infects spiroplasmas, which are known pathogens of honeybees. Here, the structure ... (SpV4) is a bacteriophage of the , which packages circular ssDNA within non-enveloped T = 1 icosahedral capsids. It infects spiroplasmas, which are known pathogens of honeybees. Here, the structure of the SpV4 virion is determined using cryo-electron microscopy to a resolution of 2.5 Å. A striking feature of the SpV4 capsid is the mushroom-like protrusions at the 3-fold axes, which is common among all members of the subfamily While the function of the protrusion is currently unknown, this feature varies widely in this subfamily and is therefore possibly an adaptation for host recognition. Furthermore, on the interior of the SpV4 capsid, the location of DNA-binding protein VP8 was identified and shown to have low structural conservation to the capsids of other viruses in the family. The structural characterization of SpV4 will aid future studies analyzing the virus-host interaction, to understand disease mechanisms at a molecular level. Furthermore, the structural comparisons in this study, including a low-resolution structure of the chlamydia phage 2, provide an overview of the structural repertoire of the viruses in this family that infect various bacterial hosts, which in turn infect a wide range of animals and plants. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_45583.map.gz emd_45583.map.gz | 589.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-45583-v30.xml emd-45583-v30.xml emd-45583.xml emd-45583.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_45583.png emd_45583.png | 58.6 KB | ||
| Filedesc metadata |  emd-45583.cif.gz emd-45583.cif.gz | 5.6 KB | ||
| Others |  emd_45583_half_map_1.map.gz emd_45583_half_map_1.map.gz emd_45583_half_map_2.map.gz emd_45583_half_map_2.map.gz | 151.5 MB 151.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-45583 http://ftp.pdbj.org/pub/emdb/structures/EMD-45583 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45583 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45583 | HTTPS FTP |
-Related structure data
| Related structure data |  9cgmMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_45583.map.gz / Format: CCP4 / Size: 634.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_45583.map.gz / Format: CCP4 / Size: 634.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.061 Å | ||||||||||||||||||||||||||||||||||||
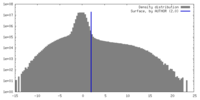
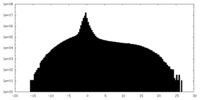
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_45583_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
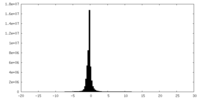
| Density Histograms |
-Half map: #2
| File | emd_45583_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
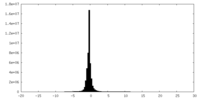
| Density Histograms |
- Sample components
Sample components
-Entire : Spiromicrovirus SpV4
| Entire | Name:  Spiromicrovirus SpV4 Spiromicrovirus SpV4 |
|---|---|
| Components |
|
-Supramolecule #1: Spiromicrovirus SpV4
| Supramolecule | Name: Spiromicrovirus SpV4 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: SpV4 was propagated in the S. melliferum strain G1 / NCBI-ID: 10855 / Sci species name: Spiromicrovirus SpV4 / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No |
|---|---|
| Virus shell | Shell ID: 1 / T number (triangulation number): 1 |
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Spiromicrovirus SpV4 Spiromicrovirus SpV4 |
| Molecular weight | Theoretical: 62.294344 KDa |
| Recombinant expression | Organism:  Spiroplasma melliferum (bacteria) Spiroplasma melliferum (bacteria) |
| Sequence | String: MKKKMSKLNA RVHDFSMFKG NHIPRSKIHI PHKTIRAFNV GEIIPIYQTP VYPGEHIKMD LTSLYRPSTF IVPPMDDLIV DTYAFAVPW RIVWKDLEKF FGENSDSWDV KNAPPVPDIV APSGGWDYGT LADHFGITPK VPGIRVKSLR FRAYAKIIND W FRDQNLSS ...String: MKKKMSKLNA RVHDFSMFKG NHIPRSKIHI PHKTIRAFNV GEIIPIYQTP VYPGEHIKMD LTSLYRPSTF IVPPMDDLIV DTYAFAVPW RIVWKDLEKF FGENSDSWDV KNAPPVPDIV APSGGWDYGT LADHFGITPK VPGIRVKSLR FRAYAKIIND W FRDQNLSS ECALTLDSSN SQGSNGSNQV TDIQLGGKPY IANKYHDYFT SCLPAPQKGA PTTLNVGGMA PVTTKFRDVP NL SGTPLIF RDNKGRTIKT GQLGIGPVDA GFLVAQNTAQ AANGERAIPS NLWADLSNAT GISISDLRLA ITYQHYKEMD ARG GTRYVE FTLNHFGVHT ADARLQRSEF LGGHSQSLLV QSVPQTSSTV EKMTPQGNLA AFSETMIQNN YLVNKTFTEH SYII VLAVV RYKHTYQQGI EADWFRGQDK FDMYDPLLAN ISEQPVKNRE IMVQGNSQDN EIFGFQEAWA DLRFKPNSVA GVMRS SHPQ SLDYWHFADH YAQLPKLSSE WLKEDYKNVD RTLALKASDN TPQLRVDFMF NTIAEKPMPL YSTPGLRRI UniProtKB: Capsid protein VP1 |
-Macromolecule #2: DNA binding protein ORF8
| Macromolecule | Name: DNA binding protein ORF8 / type: protein_or_peptide / ID: 2 / Number of copies: 60 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Spiromicrovirus SpV4 Spiromicrovirus SpV4 |
| Molecular weight | Theoretical: 4.644519 KDa |
| Recombinant expression | Organism:  Spiroplasma melliferum (bacteria) Spiroplasma melliferum (bacteria) |
| Sequence | String: MRRKVKNTKR HQWRLTHSAR SIKRANIMPS NPRGGRRF UniProtKB: DNA binding protein ORF8 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 34.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Resolution.type: BY AUTHOR / Resolution: 2.52 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 77204 |
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)