[English] 日本語
 Yorodumi
Yorodumi- EMDB-44644: SARS CoV2 spike in complex with NTD-directed antibody Fab DH1052 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS CoV2 spike in complex with NTD-directed antibody Fab DH1052 | |||||||||
 Map data Map data | CoV2 spike in complex with antibody Fab DH1052 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS coronavirus spike antibody / VIRAL PROTEIN | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
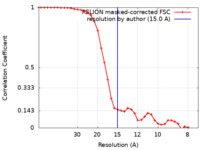
| Method | single particle reconstruction / negative staining / Resolution: 15.0 Å | |||||||||
 Authors Authors | Edwards RJ / Mansouri K | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog. Journal: PLoS Pathog.Title: Non-neutralizing SARS-CoV-2 N-terminal domain antibodies protects mice against severe disease using Fc effector functions Authors: Pierre CN / Adams LE / Anasti K / Goodman D / Stanfield-Oakley S / Li D / Rountree W / Wang Y / Edwards RJ / Alam SM / Ferrari G / Tomaras GD / Haynes BF / Baric RS / Saunders KO | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44644.map.gz emd_44644.map.gz | 3.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44644-v30.xml emd-44644-v30.xml emd-44644.xml emd-44644.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44644_fsc.xml emd_44644_fsc.xml | 3.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_44644.png emd_44644.png | 51.6 KB | ||
| Filedesc metadata |  emd-44644.cif.gz emd-44644.cif.gz | 5.1 KB | ||
| Others |  emd_44644_half_map_1.map.gz emd_44644_half_map_1.map.gz emd_44644_half_map_2.map.gz emd_44644_half_map_2.map.gz | 2.4 MB 2.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44644 http://ftp.pdbj.org/pub/emdb/structures/EMD-44644 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44644 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44644 | HTTPS FTP |
-Validation report
| Summary document |  emd_44644_validation.pdf.gz emd_44644_validation.pdf.gz | 599.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_44644_full_validation.pdf.gz emd_44644_full_validation.pdf.gz | 599.4 KB | Display | |
| Data in XML |  emd_44644_validation.xml.gz emd_44644_validation.xml.gz | 9 KB | Display | |
| Data in CIF |  emd_44644_validation.cif.gz emd_44644_validation.cif.gz | 11.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44644 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44644 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44644 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44644 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_44644.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44644.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CoV2 spike in complex with antibody Fab DH1052 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 3.74 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map 1
| File | emd_44644_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
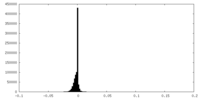
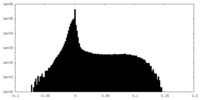
| Density Histograms |
-Half map: Half map 2
| File | emd_44644_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
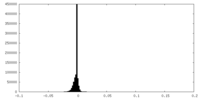
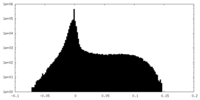
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of SARS CoV2 spike with NTD-directed antibody Fab DH1052
| Entire | Name: Complex of SARS CoV2 spike with NTD-directed antibody Fab DH1052 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of SARS CoV2 spike with NTD-directed antibody Fab DH1052
| Supramolecule | Name: Complex of SARS CoV2 spike with NTD-directed antibody Fab DH1052 type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 372 KDa |
-Supramolecule #2: DH1052 antibody Fab
| Supramolecule | Name: DH1052 antibody Fab / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: SARS CoV2 spike
| Supramolecule | Name: SARS CoV2 spike / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Staining | Type: NEGATIVE / Material: Uranyl formate Details: A 5 microliter drop of complex was incubated on the grid for 10 to 15 s, blotted and then stained with 2 percent uranyl formate for 1 minute, then blotted and air dried. | |||||||||
| Grid | Model: Homemade / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 3 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. | |||||||||
| Details | Spike protein and Fab protein at 1 mg per ml were stored at minus 80 degrees C until use. Spike was thawed in an aluminum block at 37 degrees C for 5 minutes, Fab was thawed in an aluminum block at room temperature for 5 minutes. To for the complex, 2 microliters of spike was mixed with Fab at a 1 to 9 ratio spike to Fab, and incubated at 37 degrees C for 1 hour. The complex was then crosslinked by diluting to a final spike concentration of 0.1 mg per ml into room temperature buffer containing 150 mM NaCl, 20 mM HEPES pH 7.4, 5 percent glycerol, and 7.5 mM glutaraldehyde. After 5 minutes cross-linking, excess glutaraldehyde was quenched by adding sufficient 1 M Tris pH 7.4 stock to give a final concentration of 75 mM Tris and incubated for 5 minutes. |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS EM420 |
|---|---|
| Image recording | Film or detector model: OTHER / Digitization - Dimensions - Width: 2048 pixel / Digitization - Dimensions - Height: 2048 pixel / Number grids imaged: 1 / Number real images: 200 / Average exposure time: 0.5 sec. / Average electron dose: 32.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 82000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross correlation |
 Movie
Movie Controller
Controller



 Z
Z Y
Y X
X