[English] 日本語
 Yorodumi
Yorodumi- EMDB-4443: Structure of left-handed protein cage consisting of 24 eleven-mem... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4443 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of left-handed protein cage consisting of 24 eleven-membered ring proteins held together by gold (I) bridges. | |||||||||
 Map data Map data | map of the left-handed protein cage | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | Transcription attenuation protein MtrB / Tryptophan RNA-binding attenuator protein domain / Tryptophan RNA-binding attenuator protein / Tryptophan RNA-binding attenuator protein-like domain superfamily / DNA-templated transcription termination / regulation of DNA-templated transcription / RNA binding / identical protein binding / Transcription attenuation protein MtrB Function and homology information Function and homology information | |||||||||
| Biological species |   Geobacillus stearothermophilus (bacteria) Geobacillus stearothermophilus (bacteria) | |||||||||
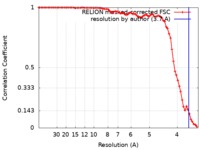
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Malay AD / Miyazaki N / Biela AP / Iwasaki K / Heddle JG | |||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: An ultra-stable gold-coordinated protein cage displaying reversible assembly. Authors: Ali D Malay / Naoyuki Miyazaki / Artur Biela / Soumyananda Chakraborti / Karolina Majsterkiewicz / Izabela Stupka / Craig S Kaplan / Agnieszka Kowalczyk / Bernard M A G Piette / Georg K A ...Authors: Ali D Malay / Naoyuki Miyazaki / Artur Biela / Soumyananda Chakraborti / Karolina Majsterkiewicz / Izabela Stupka / Craig S Kaplan / Agnieszka Kowalczyk / Bernard M A G Piette / Georg K A Hochberg / Di Wu / Tomasz P Wrobel / Adam Fineberg / Manish S Kushwah / Mitja Kelemen / Primož Vavpetič / Primož Pelicon / Philipp Kukura / Justin L P Benesch / Kenji Iwasaki / Jonathan G Heddle /       Abstract: Symmetrical protein cages have evolved to fulfil diverse roles in nature, including compartmentalization and cargo delivery, and have inspired synthetic biologists to create novel protein assemblies ...Symmetrical protein cages have evolved to fulfil diverse roles in nature, including compartmentalization and cargo delivery, and have inspired synthetic biologists to create novel protein assemblies via the precise manipulation of protein-protein interfaces. Despite the impressive array of protein cages produced in the laboratory, the design of inducible assemblies remains challenging. Here we demonstrate an ultra-stable artificial protein cage, the assembly and disassembly of which can be controlled by metal coordination at the protein-protein interfaces. The addition of a gold (I)-triphenylphosphine compound to a cysteine-substituted, 11-mer protein ring triggers supramolecular self-assembly, which generates monodisperse cage structures with masses greater than 2 MDa. The geometry of these structures is based on the Archimedean snub cube and is, to our knowledge, unprecedented. Cryo-electron microscopy confirms that the assemblies are held together by 120 S-Au-S staples between the protein oligomers, and exist in two chiral forms. The cage shows extreme chemical and thermal stability, yet it readily disassembles upon exposure to reducing agents. As well as gold, mercury(II) is also found to enable formation of the protein cage. This work establishes an approach for linking protein components into robust, higher-order structures, and expands the design space available for supramolecular assemblies to include previously unexplored geometries. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4443.map.gz emd_4443.map.gz | 9.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4443-v30.xml emd-4443-v30.xml emd-4443.xml emd-4443.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4443_fsc.xml emd_4443_fsc.xml | 7.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_4443.png emd_4443.png | 100 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4443 http://ftp.pdbj.org/pub/emdb/structures/EMD-4443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4443 | HTTPS FTP |
-Related structure data
| Related structure data |  6rvvMC  4444C  6966C  6rvwC  6962 M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4443.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4443.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map of the left-handed protein cage | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.74 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Designed protein cage consisting of C11-symmetric TRAP proteins c...
| Entire | Name: Designed protein cage consisting of C11-symmetric TRAP proteins coordinated by Au+1 ions |
|---|---|
| Components |
|
-Supramolecule #1: Designed protein cage consisting of C11-symmetric TRAP proteins c...
| Supramolecule | Name: Designed protein cage consisting of C11-symmetric TRAP proteins coordinated by Au+1 ions type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Geobacillus stearothermophilus (bacteria) Geobacillus stearothermophilus (bacteria) |
| Recombinant expression | Organism:  |
| Molecular weight | Experimental: 2.2 MDa |
-Macromolecule #1: Transcription attenuation protein MtrB
| Macromolecule | Name: Transcription attenuation protein MtrB / type: protein_or_peptide / ID: 1 / Number of copies: 264 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Geobacillus stearothermophilus (bacteria) Geobacillus stearothermophilus (bacteria) |
| Molecular weight | Theoretical: 8.161224 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MYTNSDFVVI KALEDGVNVI GLTRGADTRF HHSECLDKGE VLIAQFTEHT SAIKVRGKAY IQTSHGVIES EGKK |
-Macromolecule #2: GOLD ION
| Macromolecule | Name: GOLD ION / type: ligand / ID: 2 / Number of copies: 120 / Formula: AU |
|---|---|
| Molecular weight | Theoretical: 196.967 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.89 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: MOLYBDENUM / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 10.0 nm / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: 3.0 s blotting time. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: OTHER Target criteria: gradient-driven minimization of combined map and restraints target | ||||||||||||||||||||||||
| Output model |  PDB-6rvv: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)