[English] 日本語
 Yorodumi
Yorodumi- EMDB-44224: Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (singly Ub-loaded) - cluster 5 map and model (ATP/Mg) | |||||||||
 Map data Map data | Full map from the gold-standard refinement, globally sharpened using an B-factor of -60 A^2, used for model building and refinement. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ubiquitin / E1 / E2 / UBA1 / UBC4 / transthioesterification / thioester / transthiolation / tetrahedral intermediate / adenylation / inhibitor / LIGASE / nucleus / phosphoprotein / UBL conjugation pathway / UBL / ATP / ATP-binding / AMP / nucleotide-binding / isopeptide bond | |||||||||
| Function / homology |  Function and homology information Function and homology informationPeroxisomal protein import / : / : / Antigen processing: Ubiquitination & Proteasome degradation / SREBP signaling pathway / E1 ubiquitin-activating enzyme / ubiquitin activating enzyme activity / positive regulation of mitotic metaphase/anaphase transition / E2 ubiquitin-conjugating enzyme / SCF-dependent proteasomal ubiquitin-dependent protein catabolic process ...Peroxisomal protein import / : / : / Antigen processing: Ubiquitination & Proteasome degradation / SREBP signaling pathway / E1 ubiquitin-activating enzyme / ubiquitin activating enzyme activity / positive regulation of mitotic metaphase/anaphase transition / E2 ubiquitin-conjugating enzyme / SCF-dependent proteasomal ubiquitin-dependent protein catabolic process / ubiquitin conjugating enzyme activity / ribosomal large subunit export from nucleus / modification-dependent protein catabolic process / protein tag activity / protein polyubiquitination / ubiquitin-protein transferase activity / cytosolic large ribosomal subunit / ubiquitin-dependent protein catabolic process / cytoplasmic translation / structural constituent of ribosome / protein ubiquitination / DNA damage response / ubiquitin protein ligase binding / nucleolus / magnesium ion binding / ATP hydrolysis activity / ATP binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
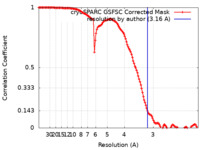
| Method | single particle reconstruction / cryo EM / Resolution: 3.16 Å | |||||||||
 Authors Authors | Kochanczyk T / Lima CD | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Structural basis for transthiolation intermediates in the ubiquitin pathway. Authors: Tomasz Kochańczyk / Zachary S Hann / Michaelyn C Lux / Avelyn Mae V Delos Reyes / Cheng Ji / Derek S Tan / Christopher D Lima /  Abstract: Transthiolation (also known as transthioesterification) reactions are used in the biosynthesis of acetyl coenzyme A, fatty acids and polyketides, and for post-translational modification by ubiquitin ...Transthiolation (also known as transthioesterification) reactions are used in the biosynthesis of acetyl coenzyme A, fatty acids and polyketides, and for post-translational modification by ubiquitin (Ub) and ubiquitin-like (Ubl) proteins. For the Ub pathway, E1 enzymes catalyse transthiolation from an E1~Ub thioester to an E2~Ub thioester. Transthiolation is also required for transfer of Ub from an E2~Ub thioester to HECT (homologous to E6AP C terminus) and RBR (ring-between-ring) E3 ligases to form E3~Ub thioesters. How isoenergetic transfer of thioester bonds is driven forward by enzymes in the Ub pathway remains unclear. Here we isolate mimics of transient transthiolation intermediates for E1-Ub(T)-E2 and E2-Ub(T)-E3 complexes (where T denotes Ub in a thioester or Ub undergoing transthiolation) using a chemical strategy with native enzymes and near-native Ub to capture and visualize a continuum of structures determined by single-particle cryo-electron microscopy. These structures and accompanying biochemical experiments illuminate conformational changes in Ub, E1, E2 and E3 that are coordinated with the chemical reactions to facilitate directional transfer of Ub from each enzyme to the next. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44224.map.gz emd_44224.map.gz | 111.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44224-v30.xml emd-44224-v30.xml emd-44224.xml emd-44224.xml | 46.3 KB 46.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44224_fsc.xml emd_44224_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_44224.png emd_44224.png | 133.7 KB | ||
| Filedesc metadata |  emd-44224.cif.gz emd-44224.cif.gz | 7.9 KB | ||
| Others |  emd_44224_additional_1.map.gz emd_44224_additional_1.map.gz emd_44224_additional_10.map.gz emd_44224_additional_10.map.gz emd_44224_additional_11.map.gz emd_44224_additional_11.map.gz emd_44224_additional_2.map.gz emd_44224_additional_2.map.gz emd_44224_additional_3.map.gz emd_44224_additional_3.map.gz emd_44224_additional_4.map.gz emd_44224_additional_4.map.gz emd_44224_additional_5.map.gz emd_44224_additional_5.map.gz emd_44224_additional_6.map.gz emd_44224_additional_6.map.gz emd_44224_additional_7.map.gz emd_44224_additional_7.map.gz emd_44224_additional_8.map.gz emd_44224_additional_8.map.gz emd_44224_additional_9.map.gz emd_44224_additional_9.map.gz emd_44224_half_map_1.map.gz emd_44224_half_map_1.map.gz emd_44224_half_map_2.map.gz emd_44224_half_map_2.map.gz | 108.7 MB 108.5 MB 108.4 MB 108.3 MB 108.3 MB 108.5 MB 108.5 MB 108.4 MB 108.4 MB 108.5 MB 108.5 MB 200.6 MB 200.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44224 http://ftp.pdbj.org/pub/emdb/structures/EMD-44224 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44224 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44224 | HTTPS FTP |
-Related structure data
| Related structure data |  9b5tMC  9b55C  9b56C  9b57C  9b58C  9b59C  9b5aC  9b5bC  9b5cC  9b5dC  9b5eC  9b5fC  9b5gC  9b5hC  9b5iC  9b5jC  9b5kC  9b5lC  9b5mC  9b5nC  9b5oC  9b5pC  9b5qC  9b5rC  9b5sC  9b5uC  9b5vC  9b5wC  9b5xC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_44224.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44224.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map from the gold-standard refinement, globally sharpened using an B-factor of -60 A^2, used for model building and refinement. | ||||||||||||||||||||||||||||||||||||
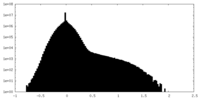
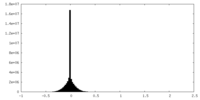
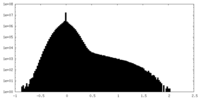
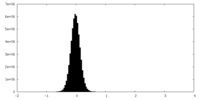
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.064 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
+Additional map: Full map from the gold-standard refinement (unsharpened).
+Additional map: Full map of Ub(T) 3D class 10 reconstructed...
+Additional map: Full map of Ub(T) 3D class 8 reconstructed...
+Additional map: Full map of Ub(T) 3D class 7 reconstructed...
+Additional map: Full map of Ub(T) 3D class 5 reconstructed...
+Additional map: Full map of Ub(T) 3D class 1 reconstructed...
+Additional map: Full map of Ub(T) 3D class 9 reconstructed...
+Additional map: Full map of Ub(T) 3D class 4 reconstructed...
+Additional map: Full map of Ub(T) 3D class 6 reconstructed...
+Additional map: Full map of Ub(T) 3D class 3 reconstructed...
+Additional map: Full map of Ub(T) 3D class 2 reconstructed...
+Half map: Half map 1 from the gold-standard refinement.
+Half map: Half map 2 from the gold-standard refinement.
- Sample components
Sample components
-Entire : Covalent E1-Ub-E2 transthiolation intermediate mimic complex (sin...
| Entire | Name: Covalent E1-Ub-E2 transthiolation intermediate mimic complex (singly Ub-loaded E1-Ub-E2 complex) |
|---|---|
| Components |
|
-Supramolecule #1: Covalent E1-Ub-E2 transthiolation intermediate mimic complex (sin...
| Supramolecule | Name: Covalent E1-Ub-E2 transthiolation intermediate mimic complex (singly Ub-loaded E1-Ub-E2 complex) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ubiquitin-activating enzyme E1 1
| Macromolecule | Name: Ubiquitin-activating enzyme E1 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: E1 ubiquitin-activating enzyme |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 111.764047 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SNTIDEGLYS RQLYVLGHEA MKQMSQSNVL IIGCKGLGVE IAKNVCLAGV KSVTLYDPQP TRIEDLSSQY FLTEDDIGVP RAKVTVSKL AELNQYVPVS VVDELSTEYL KNFKCVVVTE TSLTKQLEIN DFTHKNHIAY IAADSRGLFG SIFCDFGENF I CTDTDGNE ...String: SNTIDEGLYS RQLYVLGHEA MKQMSQSNVL IIGCKGLGVE IAKNVCLAGV KSVTLYDPQP TRIEDLSSQY FLTEDDIGVP RAKVTVSKL AELNQYVPVS VVDELSTEYL KNFKCVVVTE TSLTKQLEIN DFTHKNHIAY IAADSRGLFG SIFCDFGENF I CTDTDGNE PLTGMIASIT DDGVVTMLEE TRHGLENGDF VKFTEVKGMP GLNDGTPRKV EVKGPYTFSI GSVKDLGSAG YN GVFTQVK VPTKISFKSL RESLKDPEYV YPDFGKMMRP PQYHIAFQAL SAFADAHEGS LPRPRNDIDA AEFFEFCKKI AST LQFDVE LDEKLIKEIS YQARGDLVAM SAFLGGAVAQ EVLKATTSKF YPLKQYFYFD SLESLPSSVT ISEETCKPRG CRYD GQIAV FGSEFQEKIA SLSTFLVGAG AIGCEMLKNW AMMGVATGES GHISVTDMDS IEKSNLNRQF LFRPRDVGKL KSECA STAV SIMNPSLTGK ITSYQERVGP ESEGIFGDEF FEKLSLVTNA LDNVEARMYV DRRCVFFEKP LLESGTLGTK GNTQVV VPH LTESYGSSQD PPEKSFPICT LKNFPNRIEH TIAWARDLFE GLFKQPIDNV NMYLSSPNFL ETSLKTSSNP REVLENI RD YLVTEKPLSF EECIMWARLQ FDKFFNNNIQ QLLFNFPKDS VTSTGQPFWS GPKRAPTPLS FDIHNREHFD FIVAAASL Y AFNYGLKSET DPAIYERVLA GYNPPPFAPK SGIKIQVNEN EEAPETAANK DKQELKSIAD SLPPPSSLVG FRLTPAEFE KDDDSNHHID FITAASNLRA MNYDITPADR FKTKFVAGKI VPAMCTSTAV VSGLVCLELV KLVDGKKKIE EYKNGFFNLA IGLFTFSDP IASPKMKVNG KEIDKIWDRY NLPDCTLQEL IDYFQKEEGL EVTMLSSGVS LLYANFQPPK KLAERLPLKI S ELVEQITK KKLEPFRKHL VLEICCDDAN GEDVEVPFIC IKL UniProtKB: Ubiquitin-activating enzyme E1 1 |
-Macromolecule #2: Ubiquitin-conjugating enzyme E2 4
| Macromolecule | Name: Ubiquitin-conjugating enzyme E2 4 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: E2 ubiquitin-conjugating enzyme |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 17.043336 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MALKRINREL ADLGKDPPSS SSAGPVGDDL FHWQATIMGP ADSPYAGGVF FLSIHFPTDY PFKPPKVNFT TRIYHPNINS NGSICLDIL RDQWSPALTI SKVLLSISSL LTDPNPDDPL VPEIAHVYKT DRSRYELSAR EWTRKYAIGG LVPR UniProtKB: Ubiquitin-conjugating enzyme E2 4 |
-Macromolecule #3: Ubiquitin
| Macromolecule | Name: Ubiquitin / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.769948 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSGGMQIFVK TLTGKTITLE VESSDTIDNV KSKIQDKEGI PPDQQRLIFA GKQLEDGRTL SDYNIQKEST LHLVLRLRG UniProtKB: Ubiquitin-ribosomal protein eL40B fusion protein |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 1 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #6: 4-aminobutanenitrile
| Macromolecule | Name: 4-aminobutanenitrile / type: ligand / ID: 6 / Number of copies: 1 / Formula: A1AIV |
|---|---|
| Molecular weight | Theoretical: 84.12 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.2 Component:
Details: 20 mM Tris-HCl, 100 mM NaCl, 5 mM MgCl2, 1 mM ATP, 0.05% CHAPSO | ||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: SUPER-RESOLUTION / Average exposure time: 4.0 sec. / Average electron dose: 72.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-9b5t: |
 Movie
Movie Controller
Controller



































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)