[English] 日本語
 Yorodumi
Yorodumi- EMDB-43893: Structure of the auto-fluorescent membrane-bound red body organel... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the auto-fluorescent membrane-bound red body organelle from Nannochloropsis oceanica in situ | |||||||||
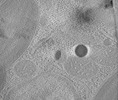
 Map data Map data | Tomographic reconstruction of a cryo-lamella through a Nannochloropsis oceanica cell showing a red body within its cellular context. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Organelle / Algae / Intracellular Transport / Carotenoid / LIPID TRANSPORT | |||||||||
| Biological species |  Nannochloropsis oceanica (eukaryote) Nannochloropsis oceanica (eukaryote) | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Grob P / Danielle J / Gee CW | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Implicating the red body of Nannochloropsis in forming the recalcitrant cell wall polymer algaenan. Authors: Christopher W Gee / Johan Andersen-Ranberg / Ethan Boynton / Rachel Z Rosen / Danielle Jorgens / Patricia Grob / Hoi-Ying N Holman / Krishna K Niyogi /   Abstract: Stramenopile algae contribute significantly to global primary productivity, and one class, Eustigmatophyceae, is increasingly studied for applications in high-value lipid production. Yet much about ...Stramenopile algae contribute significantly to global primary productivity, and one class, Eustigmatophyceae, is increasingly studied for applications in high-value lipid production. Yet much about their basic biology remains unknown, including the nature of an enigmatic, pigmented globule found in vegetative cells. Here, we present an in-depth examination of this "red body," focusing on Nannochloropsis oceanica. During the cell cycle, the red body forms adjacent to the plastid, but unexpectedly it is secreted and released with the autosporangial wall following cell division. Shed red bodies contain antioxidant ketocarotenoids, and overexpression of a beta-carotene ketolase results in enlarged red bodies. Infrared spectroscopy indicates long-chain, aliphatic lipids in shed red bodies and cell walls, and UHPLC-HRMS detects a C32 alkyl diol, a potential precursor of algaenan, a recalcitrant cell wall polymer. We propose that the red body transports algaenan precursors from plastid to apoplast to be incorporated into daughter cell walls. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43893.map.gz emd_43893.map.gz | 514.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43893-v30.xml emd-43893-v30.xml emd-43893.xml emd-43893.xml | 12.7 KB 12.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_43893.png emd_43893.png | 199.2 KB | ||
| Filedesc metadata |  emd-43893.cif.gz emd-43893.cif.gz | 5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43893 http://ftp.pdbj.org/pub/emdb/structures/EMD-43893 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43893 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43893 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_43893.map.gz / Format: CCP4 / Size: 558.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43893.map.gz / Format: CCP4 / Size: 558.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Tomographic reconstruction of a cryo-lamella through a Nannochloropsis oceanica cell showing a red body within its cellular context. | ||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 22.13 Å | ||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Nannochloropsis oceanica CCMP1779
| Entire | Name: Nannochloropsis oceanica CCMP1779 |
|---|---|
| Components |
|
-Supramolecule #1: Nannochloropsis oceanica CCMP1779
| Supramolecule | Name: Nannochloropsis oceanica CCMP1779 / type: cell / ID: 1 / Parent: 0 Details: Tomographic reconstruction of a red body organelle from Nannochloropsis oceanica in situ |
|---|---|
| Source (natural) | Organism:  Nannochloropsis oceanica (eukaryote) / Strain: CCMP1779 Nannochloropsis oceanica (eukaryote) / Strain: CCMP1779 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 8.1 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Support film - Film thickness: 12 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.027 kPa / Details: the grid was soaked in chloroform before use | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV / Details: Blotted manually from opposite side of the grid. | ||||||||||||
| Details | Nannochloropsis oceanica CCMP1779 cells were grown in artificial seawater and f-media enrichment entrained to a 12-12 light-dark photoperiod and sampled in mid-log phase (~1x10^7 cells/ml), shortly after subjective dark when cells are undergoing division. | ||||||||||||
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 / Focused ion beam - Current: 0.037 / Focused ion beam - Duration: 600 / Focused ion beam - Temperature: 123 K / Focused ion beam - Initial thickness: 1000 / Focused ion beam - Final thickness: 380 Focused ion beam - Details: Grids were transfered to a Leica Ace 900 (Leica Microsystems) for coating with 5nm of platinum prior to being transferred to the Zeiss Crossbeam 540 (Zeiss, Germany). The ...Focused ion beam - Details: Grids were transfered to a Leica Ace 900 (Leica Microsystems) for coating with 5nm of platinum prior to being transferred to the Zeiss Crossbeam 540 (Zeiss, Germany). The Zeiss Crossbeam 540 operated with a Leica CryoStage (Leica Microsystems, GmbH) cooled to -150 deg. C was used for milling and imaging of the frozen cells. To create lamellae used for cryo-tomography, milling was performed with a gallium ion source at an energy of 37 pA and a working distance of 5mm. Imaging of the grid and milled lamellae was done using an Everhart-Thornley detector at 2.0 kV.. The value given for _em_focused_ion_beam.instrument is Zeiss Crossbeam 540. This is not in a list of allowed values {'OTHER', 'DB235'} so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 35 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 100.0 e/Å2 Details: images were collected in movie mode and super resolution |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 6.0 µm / Nominal defocus min: 1.6 µm / Nominal magnification: 15000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | the movie frames were aligned with MotionCor2 and binned to normal resolution |
|---|---|
| Final reconstruction | Algorithm: BACK PROJECTION / Software - Name:  IMOD (ver. 4.11.24) / Software - details: Etomo / Number images used: 61 IMOD (ver. 4.11.24) / Software - details: Etomo / Number images used: 61 |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
















