+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human EBP complexed with compound 1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Emopamil-Binding Protein Isomerization Protein structure complex / STRUCTURAL PROTEIN / ISOMERASE-INHIBITOR complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationcholestenol Delta-isomerase / C-8 sterol isomerase activity / cholesterol biosynthetic process via desmosterol / cholesterol biosynthetic process via lathosterol / cholestenol delta-isomerase activity / Cholesterol biosynthesis via desmosterol / Cholesterol biosynthesis via lathosterol / cholesterol-5,6-oxide hydrolase / cholesterol-5,6-oxide hydrolase activity / steroid Delta-isomerase activity ...cholestenol Delta-isomerase / C-8 sterol isomerase activity / cholesterol biosynthetic process via desmosterol / cholesterol biosynthetic process via lathosterol / cholestenol delta-isomerase activity / Cholesterol biosynthesis via desmosterol / Cholesterol biosynthesis via lathosterol / cholesterol-5,6-oxide hydrolase / cholesterol-5,6-oxide hydrolase activity / steroid Delta-isomerase activity / ossification involved in bone maturation / cholesterol biosynthetic process / hemopoiesis / cholesterol metabolic process / nuclear envelope / cytoplasmic vesicle / nuclear membrane / endoplasmic reticulum membrane / endoplasmic reticulum / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
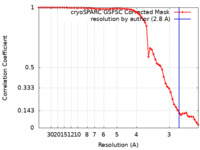
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Sun D / Masureel M | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: J Med Chem / Year: 2024 Journal: J Med Chem / Year: 2024Title: Discovery and Optimization of Selective Brain-Penetrant EBP Inhibitors that Enhance Oligodendrocyte Formation. Authors: Ruth Dorel / Dawei Sun / Nicholas Carruthers / Georgette M Castanedo / Peter M-U Ung / Daniel C Factor / Tianbo Li / Hannah Baumann / Danielle Janota / Jodie Pang / Laurent Salphati / Robert ...Authors: Ruth Dorel / Dawei Sun / Nicholas Carruthers / Georgette M Castanedo / Peter M-U Ung / Daniel C Factor / Tianbo Li / Hannah Baumann / Danielle Janota / Jodie Pang / Laurent Salphati / Robert Meklemburg / Allison J Korman / Halie E Harper / Samantha Stubblefield / Jian Payandeh / Daniel McHugh / Bradley T Lang / Paul J Tesar / Edward Dere / Matthieu Masureel / Drew J Adams / Matthew Volgraf / Marie-Gabrielle Braun /  Abstract: The inhibition of emopamil binding protein (EBP), a sterol isomerase within the cholesterol biosynthesis pathway, promotes oligodendrocyte formation, which has been proposed as a potential ...The inhibition of emopamil binding protein (EBP), a sterol isomerase within the cholesterol biosynthesis pathway, promotes oligodendrocyte formation, which has been proposed as a potential therapeutic approach for treating multiple sclerosis. Herein, we describe the discovery and optimization of brain-penetrant, orally bioavailable inhibitors of EBP. A structure-based drug design approach from literature compound led to the discovery of a hydantoin-based scaffold, which provided balanced physicochemical properties and potency and an improved safety profile. The long half-lives of early hydantoin-based EBP inhibitors in rodents prompted an unconventional optimization strategy, focused on increasing metabolic turnover while maintaining potency and a brain-penetrant profile. The resulting EBP inhibitor demonstrated strong target engagement in the brain, as illustrated by the accumulation of EBP substrate zymostenol after repeated dosing. Furthermore, compound enhanced the formation of oligodendrocytes in human cortical organoids, providing additional support for our therapeutic hypothesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43712.map.gz emd_43712.map.gz | 25.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43712-v30.xml emd-43712-v30.xml emd-43712.xml emd-43712.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43712_fsc.xml emd_43712_fsc.xml | 6.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_43712.png emd_43712.png | 104.7 KB | ||
| Filedesc metadata |  emd-43712.cif.gz emd-43712.cif.gz | 5.6 KB | ||
| Others |  emd_43712_additional_1.map.gz emd_43712_additional_1.map.gz emd_43712_half_map_1.map.gz emd_43712_half_map_1.map.gz emd_43712_half_map_2.map.gz emd_43712_half_map_2.map.gz | 13.7 MB 25 MB 25 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43712 http://ftp.pdbj.org/pub/emdb/structures/EMD-43712 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43712 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43712 | HTTPS FTP |
-Validation report
| Summary document |  emd_43712_validation.pdf.gz emd_43712_validation.pdf.gz | 845.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43712_full_validation.pdf.gz emd_43712_full_validation.pdf.gz | 844.9 KB | Display | |
| Data in XML |  emd_43712_validation.xml.gz emd_43712_validation.xml.gz | 13.8 KB | Display | |
| Data in CIF |  emd_43712_validation.cif.gz emd_43712_validation.cif.gz | 17.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43712 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43712 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43712 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43712 | HTTPS FTP |
-Related structure data
| Related structure data |  8w0rMC  8w0sC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_43712.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43712.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.2 Å | ||||||||||||||||||||||||||||||||||||
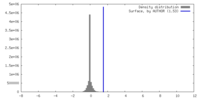
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: unsharpened map
| File | emd_43712_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
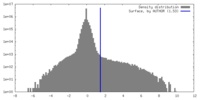
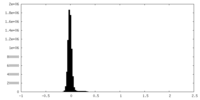
| Density Histograms |
-Half map: #2
| File | emd_43712_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
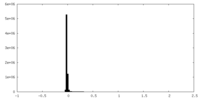
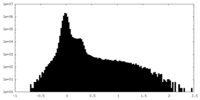
| Density Histograms |
-Half map: #1
| File | emd_43712_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
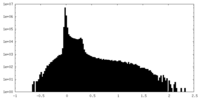
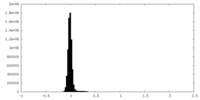
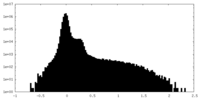
| Density Histograms |
- Sample components
Sample components
-Entire : EBP complexed with compound 1
| Entire | Name: EBP complexed with compound 1 |
|---|---|
| Components |
|
-Supramolecule #1: EBP complexed with compound 1
| Supramolecule | Name: EBP complexed with compound 1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: 3-beta-hydroxysteroid-Delta(8),Delta(7)-isomerase
| Macromolecule | Name: 3-beta-hydroxysteroid-Delta(8),Delta(7)-isomerase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: cholestenol Delta-isomerase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.421871 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MWSHPQFEKT TNAGPLHPYW PQHLRLDNFV PNDRPTWHIL AGLFSVTGVL VVTTWLLSGR AAVVPLGTWR RLSLCWFAVC GFIHLVIEG WFVLYYEDLL GDQAFLSQLW KEYAKGDSRY ILGDNFTVCM ETITACLWGP LSLWVVIAFL RQHPLRFILQ L VVSVGQIY ...String: MWSHPQFEKT TNAGPLHPYW PQHLRLDNFV PNDRPTWHIL AGLFSVTGVL VVTTWLLSGR AAVVPLGTWR RLSLCWFAVC GFIHLVIEG WFVLYYEDLL GDQAFLSQLW KEYAKGDSRY ILGDNFTVCM ETITACLWGP LSLWVVIAFL RQHPLRFILQ L VVSVGQIY GDVLYFLTEH RDGFQHGELG HPLYFWFYFV FMNALWLVLP GVLVLDAVKH LTHAQSTLDA KATKAKSKKN UniProtKB: 3-beta-hydroxysteroid-Delta(8),Delta(7)-isomerase |
-Macromolecule #2: 1-methyl-1'-[(oxan-4-yl)methyl]-5-(trifluoromethyl)spiro[indole-2...
| Macromolecule | Name: 1-methyl-1'-[(oxan-4-yl)methyl]-5-(trifluoromethyl)spiro[indole-2,4'-piperidin]-3(1H)-one type: ligand / ID: 2 / Number of copies: 2 / Formula: A1AEU |
|---|---|
| Molecular weight | Theoretical: 382.42 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 64.009 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)