[English] 日本語
 Yorodumi
Yorodumi- EMDB-43137: SARS-CoV-2 Frameshift Stimulatory Element with Upstream Multibran... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 Frameshift Stimulatory Element with Upstream Multibranch Loop | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA / Coronavirus / SARS-CoV-2 / Programmed -1 Ribosomal Frameshifting / -1 PRF / Frameshift Stimulatory Element / FSE / Attenuator Hairpin | |||||||||
| Biological species |   Severe acute respiratory syndrome coronavirus Severe acute respiratory syndrome coronavirus | |||||||||
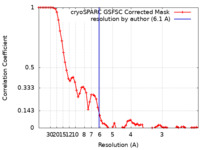
| Method | single particle reconstruction / cryo EM / Resolution: 6.1 Å | |||||||||
 Authors Authors | Peterson JM / Becker ST / O'Leary CA / Juneja P / Yang Y / Moss WN | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Biochemistry / Year: 2024 Journal: Biochemistry / Year: 2024Title: Structure of the SARS-CoV-2 Frameshift Stimulatory Element with an Upstream Multibranch Loop. Authors: Jake M Peterson / Scott T Becker / Collin A O'Leary / Puneet Juneja / Yang Yang / Walter N Moss /  Abstract: The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) frameshift stimulatory element (FSE) is necessary for programmed -1 ribosomal frameshifting (-1 PRF) and optimized viral efficacy. The ...The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) frameshift stimulatory element (FSE) is necessary for programmed -1 ribosomal frameshifting (-1 PRF) and optimized viral efficacy. The FSE has an abundance of context-dependent alternate conformations, but two of the structures most crucial to -1 PRF are an attenuator hairpin and a three-stem H-type pseudoknot structure. A crystal structure of the pseudoknot alone features three RNA stems in a helically stacked linear structure, whereas a 6.9 Å cryo-EM structure including the upstream heptameric slippery site resulted in a bend between two stems. Our previous research alluded to an extended upstream multibranch loop that includes both the attenuator hairpin and the slippery site-a conformation not previously modeled. We aim to provide further context to the SARS-CoV-2 FSE via computational and medium resolution cryo-EM approaches, by presenting a 6.1 Å cryo-EM structure featuring a linear pseudoknot structure and a dynamic upstream multibranch loop. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43137.map.gz emd_43137.map.gz | 15.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43137-v30.xml emd-43137-v30.xml emd-43137.xml emd-43137.xml | 17.9 KB 17.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43137_fsc.xml emd_43137_fsc.xml | 6.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_43137.png emd_43137.png | 132.1 KB | ||
| Masks |  emd_43137_msk_1.map emd_43137_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-43137.cif.gz emd-43137.cif.gz | 5 KB | ||
| Others |  emd_43137_additional_1.map.gz emd_43137_additional_1.map.gz emd_43137_half_map_1.map.gz emd_43137_half_map_1.map.gz emd_43137_half_map_2.map.gz emd_43137_half_map_2.map.gz | 28.6 MB 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43137 http://ftp.pdbj.org/pub/emdb/structures/EMD-43137 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43137 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43137 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_43137.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43137.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.133 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_43137_msk_1.map emd_43137_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_43137_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_43137_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_43137_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Severe acute respiratory syndrome coronavirus
| Entire | Name:  Severe acute respiratory syndrome coronavirus Severe acute respiratory syndrome coronavirus |
|---|---|
| Components |
|
-Supramolecule #1: Severe acute respiratory syndrome coronavirus
| Supramolecule | Name: Severe acute respiratory syndrome coronavirus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: PCR amplified and transcribed using T7 polymerase. / NCBI-ID: 2901879 Sci species name: Severe acute respiratory syndrome coronavirus Sci species strain: Wuhan-Hu-1 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / Strain: Wuhan-Hu-1 Homo sapiens (human) / Strain: Wuhan-Hu-1 |
-Macromolecule #1: Frameshift Stimulatory Element with Upstream Multi-branch Loop
| Macromolecule | Name: Frameshift Stimulatory Element with Upstream Multi-branch Loop type: rna / ID: 1 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 37.90141 KDa |
| Sequence | String: GGGAACCCAU GCUUCAGUCA GCUGAUGCAC AAUCGUUUUU AAACGGGUUU CCGGUGUAAG UGCAGCCCGU CUUACACCGU GCGGCACAG GCACUAGUAC UGAUGUCGUA UACAGGGCU GENBANK: GENBANK: NC_045512.2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 4 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 3 / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 36000 |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8vci: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)