[English] 日本語
 Yorodumi
Yorodumi- EMDB-42783: Triplet microtubule from the proximal region of basal body isolat... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Triplet microtubule from the proximal region of basal body isolated from a Tetrahymena thermophila strain with POC1 knockout | |||||||||
 Map data Map data | Triplet microtubule from the proximal region of basal body, poc1KO, Tetrahymena thermophila | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cilia / basal body / microtubule / CYTOSOLIC PROTEIN | |||||||||
| Biological species |  | |||||||||
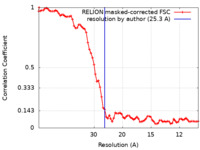
| Method | subtomogram averaging / cryo EM / Resolution: 25.3 Å | |||||||||
 Authors Authors | Ruehle M / Li S / Agard DA / Pearson C | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: J Cell Biol / Year: 2024 Journal: J Cell Biol / Year: 2024Title: Poc1 bridges basal body inner junctions to promote triplet microtubule integrity and connections. Authors: Marisa D Ruehle / Sam Li / David A Agard / Chad G Pearson /  Abstract: Basal bodies (BBs) are conserved eukaryotic structures that organize cilia. They are comprised of nine, cylindrically arranged, triplet microtubules (TMTs) connected to each other by inter-TMT ...Basal bodies (BBs) are conserved eukaryotic structures that organize cilia. They are comprised of nine, cylindrically arranged, triplet microtubules (TMTs) connected to each other by inter-TMT linkages which stabilize the structure. Poc1 is a conserved protein important for BB structural integrity in the face of ciliary forces transmitted to BBs. To understand how Poc1 confers BB stability, we identified the precise position of Poc1 in the Tetrahymena BB and the effect of Poc1 loss on BB structure. Poc1 binds at the TMT inner junctions, stabilizing TMTs directly. From this location, Poc1 also stabilizes inter-TMT linkages throughout the BB, including the cartwheel pinhead and the inner scaffold. The full localization of the inner scaffold protein Fam161A requires Poc1. As ciliary forces are increased, Fam161A is reduced, indicative of a force-dependent molecular remodeling of the inner scaffold. Thus, while not essential for BB assembly, Poc1 promotes BB interconnections that establish an architecture competent to resist ciliary forces. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42783.map.gz emd_42783.map.gz | 33.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42783-v30.xml emd-42783-v30.xml emd-42783.xml emd-42783.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42783_fsc.xml emd_42783_fsc.xml | 7.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_42783.png emd_42783.png | 58.2 KB | ||
| Masks |  emd_42783_msk_1.map emd_42783_msk_1.map | 35.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-42783.cif.gz emd-42783.cif.gz | 4.4 KB | ||
| Others |  emd_42783_half_map_1.map.gz emd_42783_half_map_1.map.gz emd_42783_half_map_2.map.gz emd_42783_half_map_2.map.gz | 27.3 MB 27.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42783 http://ftp.pdbj.org/pub/emdb/structures/EMD-42783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42783 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42783.map.gz / Format: CCP4 / Size: 35.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42783.map.gz / Format: CCP4 / Size: 35.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Triplet microtubule from the proximal region of basal body, poc1KO, Tetrahymena thermophila | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.3 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_42783_msk_1.map emd_42783_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_42783_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_42783_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Triplet microtubule from basal body isolated from Tetrahymena the...
| Entire | Name: Triplet microtubule from basal body isolated from Tetrahymena thermophila |
|---|---|
| Components |
|
-Supramolecule #1: Triplet microtubule from basal body isolated from Tetrahymena the...
| Supramolecule | Name: Triplet microtubule from basal body isolated from Tetrahymena thermophila type: organelle_or_cellular_component / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 1xTE |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK III |
| Details | Basal bodies are isolated from mutant strain POC1 knockout. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 78.0 K / Max: 78.0 K |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 6000 pixel / Digitization - Dimensions - Height: 4000 pixel / Number real images: 1 / Average exposure time: 0.36 sec. / Average electron dose: 1.33 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 18868 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 33000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)