[English] 日本語
 Yorodumi
Yorodumi- EMDB-42680: Cryo-EM structure of an Enterobacter GH43 Beta-Xylosidase: EcXyl43 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of an Enterobacter GH43 Beta-Xylosidase: EcXyl43 | |||||||||
 Map data Map data | deepenhancer map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GH43 / Enterobacter / Beta-Xylosidase / EcXyl43 / HYDROLASE | |||||||||
| Function / homology | Beta-xylosidase, C-terminal Concanavalin A-like domain / Beta xylosidase C-terminal Concanavalin A-like domain / Glycoside hydrolase, family 43 / Glycosyl hydrolases family 43 / Glycosyl hydrolase, five-bladed beta-propellor domain superfamily / hydrolase activity, hydrolyzing O-glycosyl compounds / Concanavalin A-like lectin/glucanase domain superfamily / carbohydrate metabolic process / Beta-xylosidase Function and homology information Function and homology information | |||||||||
| Biological species |  Enterobacter hormaechei (CDC Enteric Group 75) Enterobacter hormaechei (CDC Enteric Group 75) | |||||||||
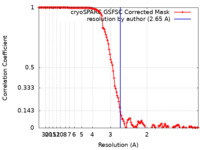
| Method | single particle reconstruction / cryo EM / Resolution: 2.65 Å | |||||||||
 Authors Authors | Briganti L / Godoy AS / Capetti CCM / Portugal RV / Polikarpov I | |||||||||
| Funding support |  Brazil, 1 items Brazil, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of an Enterobacter GH43 Beta-Xylosidase: EcXyl43 Authors: Briganti L / Godoy AS / Polikarpov I | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42680.map.gz emd_42680.map.gz | 203.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42680-v30.xml emd-42680-v30.xml emd-42680.xml emd-42680.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42680_fsc.xml emd_42680_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_42680.png emd_42680.png | 76.4 KB | ||
| Filedesc metadata |  emd-42680.cif.gz emd-42680.cif.gz | 5.9 KB | ||
| Others |  emd_42680_additional_1.map.gz emd_42680_additional_1.map.gz emd_42680_additional_2.map.gz emd_42680_additional_2.map.gz emd_42680_half_map_1.map.gz emd_42680_half_map_1.map.gz emd_42680_half_map_2.map.gz emd_42680_half_map_2.map.gz | 121 MB 1.7 MB 226.6 MB 226.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42680 http://ftp.pdbj.org/pub/emdb/structures/EMD-42680 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42680 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42680 | HTTPS FTP |
-Related structure data
| Related structure data |  8uwsMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42680.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42680.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | deepenhancer map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.67 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: cryosparc refinement map
| File | emd_42680_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryosparc refinement map | ||||||||||||
| Projections & Slices |
| ||||||||||||
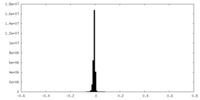
| Density Histograms |
-Additional map: mask
| File | emd_42680_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | mask | ||||||||||||
| Projections & Slices |
| ||||||||||||
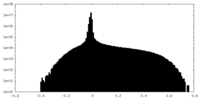
| Density Histograms |
-Half map: half B
| File | emd_42680_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half B | ||||||||||||
| Projections & Slices |
| ||||||||||||
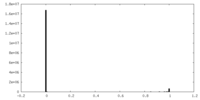
| Density Histograms |
-Half map: half A
| File | emd_42680_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half A | ||||||||||||
| Projections & Slices |
| ||||||||||||
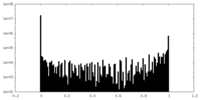
| Density Histograms |
- Sample components
Sample components
-Entire : Tetrameric GH43 structure
| Entire | Name: Tetrameric GH43 structure |
|---|---|
| Components |
|
-Supramolecule #1: Tetrameric GH43 structure
| Supramolecule | Name: Tetrameric GH43 structure / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Enterobacter hormaechei (CDC Enteric Group 75) Enterobacter hormaechei (CDC Enteric Group 75) |
-Macromolecule #1: Beta-xylosidase
| Macromolecule | Name: Beta-xylosidase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Enterobacter hormaechei (CDC Enteric Group 75) Enterobacter hormaechei (CDC Enteric Group 75) |
| Molecular weight | Theoretical: 61.000953 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEITNPILTG FNPDPSLCRQ GEDYYIATST FEWFPGVRIY HSRDLKNWTL VSTPLDRVSM LDMKGNPDSG GIWAPCLSYA DGKFWLLYT DVKIVDSPWK NGRNFLVTAP SIEGPWSEPI PMGNGGFDPS LFHDDDGRKY YLYRPWGPRH HSNPHNTIVM Q EFDPQTGT ...String: MEITNPILTG FNPDPSLCRQ GEDYYIATST FEWFPGVRIY HSRDLKNWTL VSTPLDRVSM LDMKGNPDSG GIWAPCLSYA DGKFWLLYT DVKIVDSPWK NGRNFLVTAP SIEGPWSEPI PMGNGGFDPS LFHDDDGRKY YLYRPWGPRH HSNPHNTIVM Q EFDPQTGT LSPERKTLFT GTPLCYTEGA HLYRHAGWYY LMVAEGGTSY EHAVVVLRAK TIDGPYELHP DVTMMTSWHL PE NPLQKSG HGSLLQTHTG EWYMAYLTSR PLRLPGVPLL ASGGRGYCPL GRETGIARIE WRDGWPYVEG GKHAQLTVKG PQV AEQPAA VQGSWRDDFD GSTLDPELQT LRIPFDDTLG SLTARPGYLR LYGNDSLNST FTQSTVARRW QHFIFRAETR MQFS PVHFQ QSAGLTCYYN SKNWSYCFVD YEEGQGRTIK VIQLDHNVPS WPLHEQPIPV PEQAESVWLR VDVDRLVYRY SYSFD GETW HAVPVTYEAW KLSDDYIGGR GFFTGAFVGL HCEDISGDGC HADFDYFTYE PA UniProtKB: Beta-xylosidase |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)