+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of the Helicobacter pylori cagYdAP PR | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | T4SS / protein translocation / MEMBRANE PROTEIN | ||||||||||||
| Biological species |  Helicobacter pylori 26695 (bacteria) Helicobacter pylori 26695 (bacteria) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.6 Å | ||||||||||||
 Authors Authors | Roberts JR | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Life Science Alliance / Year: 2024 Journal: Life Science Alliance / Year: 2024Title: Subdomains of the H. pylori Cag T4SS outer membrane core complex exhibit structural independence Authors: Roberts JR / Tran SC / Frick-Cheng AF / Bryant KN / Okoye CD / Cover TL / Ohi MD | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42392.map.gz emd_42392.map.gz | 978.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42392-v30.xml emd-42392-v30.xml emd-42392.xml emd-42392.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
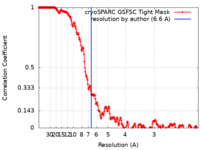
| FSC (resolution estimation) |  emd_42392_fsc.xml emd_42392_fsc.xml | 21.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_42392.png emd_42392.png | 69.1 KB | ||
| Filedesc metadata |  emd-42392.cif.gz emd-42392.cif.gz | 5.6 KB | ||
| Others |  emd_42392_half_map_1.map.gz emd_42392_half_map_1.map.gz emd_42392_half_map_2.map.gz emd_42392_half_map_2.map.gz | 964.5 MB 964.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42392 http://ftp.pdbj.org/pub/emdb/structures/EMD-42392 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42392 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42392 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_42392.map.gz / Format: CCP4 / Size: 1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42392.map.gz / Format: CCP4 / Size: 1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_42392_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_42392_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Helicobacter pylori CagYdAP PR
| Entire | Name: Helicobacter pylori CagYdAP PR |
|---|---|
| Components |
|
-Supramolecule #1: Helicobacter pylori CagYdAP PR
| Supramolecule | Name: Helicobacter pylori CagYdAP PR / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Helicobacter pylori 26695 (bacteria) / Strain: 26695 / Location in cell: membrane Helicobacter pylori 26695 (bacteria) / Strain: 26695 / Location in cell: membrane |
-Macromolecule #1: Cag pathogenicity island protein (Cag8)
| Macromolecule | Name: Cag pathogenicity island protein (Cag8) / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Helicobacter pylori 26695 (bacteria) Helicobacter pylori 26695 (bacteria) |
| Sequence | String: MGQAFFKKIV GCFCLGYLF L SSAIEAAA LD IKNFNRG RVK VVNKKI AYLG DEKPI TIWTS LDNV TVIQLE KDE TISYITT GF NKGWSIVP N SNHIFIQPK SVKSNLMFEK EAVNFALMT R DYQEFLKT KK LIVDAPD PKE LEEQKK ALEK EKEAK ...String: MGQAFFKKIV GCFCLGYLF L SSAIEAAA LD IKNFNRG RVK VVNKKI AYLG DEKPI TIWTS LDNV TVIQLE KDE TISYITT GF NKGWSIVP N SNHIFIQPK SVKSNLMFEK EAVNFALMT R DYQEFLKT KK LIVDAPD PKE LEEQKK ALEK EKEAK EQAQK AQKD KREKRK EER AKNRANL EN LTNAMSNP Q NLSNNKNLS EFIKQQRENE LDQMERLED M QEQAQANA LK QIEELNK KQA EETIKQ RAKD KINIK TDKPQ KSPE DNSIEL SPS DSAWRTN LV VRTNKALY Q FILRIAQKD NFASAYLTVK LEYPQRHEV S SVIEELKK RE EAKRQKE LIK QENLNT TAYI NRVMM ASNEQ IINK EKIREE KQK IILDQAK AL ETQYVHNA L KRNPVPRNY NYYQAPEKRS KHIMPSEIF D DGTFTYFG FK NITLQPA IFV VQPDGK LSMT DAAID PNMTN SGLR WYRVNE IAE KFKLIKD KA LVTVINKG Y GKNPLTKNY NIKNYGELER VIKKLPEVR D K |
-Macromolecule #2: Cag pathogenicity island protein (Cag7)
| Macromolecule | Name: Cag pathogenicity island protein (Cag7) / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Helicobacter pylori 26695 (bacteria) Helicobacter pylori 26695 (bacteria) |
| Sequence | String: MNEENDKLET SKKAQQDSPQ DLSNEEATEA NHFENLLKES KESSDHHLDN PTETQTHFDG DKSEETQTQM DSEGNETSES SNGSLADKLF KKARKLVDNK KPFTQQKNLD EETQELNEED DQENNEYQEE TQTDLIDDET SKKTQQHSPQ DLSNEEATEA NHFENLLKES ...String: MNEENDKLET SKKAQQDSPQ DLSNEEATEA NHFENLLKES KESSDHHLDN PTETQTHFDG DKSEETQTQM DSEGNETSES SNGSLADKLF KKARKLVDNK KPFTQQKNLD EETQELNEED DQENNEYQEE TQTDLIDDET SKKTQQHSPQ DLSNEEATEA NHFENLLKES KESSDHHLDN PTETQTNFDG DKSEETQTQM DSEGNETSES SNGSLADKLF KKARKLVDNK KPFTQQKNLD EETQELNEED DQENNEYQEE TQTDLIDDET SKKTQQHSPQ DLSNEEATEA NHFENLLKES KESSDHHLDN PTETQTNFDG DKSEEITDDS NDQEIIKGSK KKYIIGGIVV AVLIVIILFS RSIFHYFMPL EDKSSRFSKD RNLYVNDEIQ IRQEYNRLLK ERNEKGNMID KNLFFNDDPN RTLYNYLNIA EIEDKNPLRA FYECISNGGN YEECLKLIKD KKLQDQMKKT LEAYNDCIKN AKTEEERIKC LDLIKDENLK KSLLNQQKVQ VALDCLKNAK TDEERNECLK LINDPEIREK FRKELELQKE LQEYKDCIKN AKTEAEKNKC LKGLSKEAIE RLKQQALDCL KNAKTDEERN ECLKNIPQDL QKELLADMSV KAYKDCVSKA RNEKEKQECE KLLTPEARKK LEQQVLDCLK NAKTDEERKK CLKDLPKDLQ SDILAKESLK AYKDCVSQAK TEAEKKECEK LLTPEAKKLL EEEAKESVKA YLDCVSQAKT EAEKKECEKL LTPEAKKKLE EAKKSVKAYL DCVSRARNEK EKKECEKLLT PEAKKLLEQQ ALDCLKNAKT DKERKKCLKD LPKDLQKKVL AKESVKAYLD CVSQAKTEAE KKECEKLLTP EARKLLEEAK KSVKAYLDCV SQAKTEAEKK ECEKLLTPEA RKLLEEXAKE SVKAYLDCVS QAKNEAEKKE CEKLLTLESK KKLEEAKKSV KAYLDCVSQA KTEAEKKECE KLLTPEAKKL LEQQALDCLK NAKTEADKKR CVKDLPKDLQ KKVLAKESLK AYKDCVSKAR NEKEKKECEK LLTPEAKKLL EEAKKSVKAY LDCVSQAKTE AEKKECEKLL TPEARKLLEE AKESVKAYKD CVSKARNEKE KKECEKLLTP EAKKLLEQQV LDCLKNAKTE ADKKRCVKDL PKDLQKKVLA KESVKAYLDC VSRARNEKEK KECEKLLTPE AKKLLEEAKE SLKAYKDCLS QARNEEERRA CEKLLTPEAR KLLEQEVKKS IKAYLDCVSR ARNEKEKKEC EKLLTPEARK FLAKQVLNCL EKAGNEEERK ACLKNLPKDL QENILAKESL KAYKDCLSQA RNEEERRACE KLLTPEARKL LEQEVKKSVK AYLDCVSRAR NEKEKKECEK LLTPEARKFL AKELQQKDKA IKDCLKNADP NDRAAIMKCL DGLSDEEKLK YLQEAREKAV ADCLAMAKTD EEKRKCQNLY SDLIQEIQNK RTQNKQNQLS KTERLHQASE CLDNLDDPTD QEAIEQCLEG LSDSERALIL GIKRQADEVD LIYSDLRNRK TFDNMAAKGY PLLPMDFKNG GDIATINATN VDADKIASDN PIYASIEPDI AKQYETEKTI KDKNLEAKLA KALGGNKKDD DKEKSKKSTA EAKAENNKID KDVAETAKNI SEIALKNKKE KSGEFVDENG NPIDDKKKAE KQDETSPVKQ AFIGKSDPTF VLAQYTPIEI TLTSKVDATL TGIVSGVVAK DVWNMNGTMI LLDKGTKVYG NYQSVKGGTP IMTRLMIVFT KAITPDGVII PLANAQAAGM LGEAGVDGYV NNMNIPPSFY KNEGDSIKIL TMDDIDFSGV YDVKITNKSV VDEIIKQSTK TLSREHEEIT TSPKGGN |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)